A compendium of kinetic modulatory profiles identifies ferroptosis regulators
- PMID: 33686292
- PMCID: PMC8159879
- DOI: 10.1038/s41589-021-00751-4
A compendium of kinetic modulatory profiles identifies ferroptosis regulators
Abstract
Cell death can be executed by regulated apoptotic and nonapoptotic pathways, including the iron-dependent process of ferroptosis. Small molecules are essential tools for studying the regulation of cell death. Using time-lapse imaging and a library of 1,833 bioactive compounds, we assembled a large compendium of kinetic cell death modulatory profiles for inducers of apoptosis and ferroptosis. From this dataset we identify dozens of ferroptosis suppressors, including numerous compounds that appear to act via cryptic off-target antioxidant or iron chelating activities. We show that the FDA-approved drug bazedoxifene acts as a potent radical trapping antioxidant inhibitor of ferroptosis both in vitro and in vivo. ATP-competitive mechanistic target of rapamycin (mTOR) inhibitors, by contrast, are on-target ferroptosis inhibitors. Further investigation revealed both mTOR-dependent and mTOR-independent mechanisms that link amino acid metabolism to ferroptosis sensitivity. These results highlight kinetic modulatory profiling as a useful tool to investigate cell death regulation.
Conflict of interest statement
Competing financial interest
S.J.D. is a member of the scientific advisory board for Ferro Therapeutics, has consulted for Toray Industries and AbbVie Inc., and is an inventor on patents related to ferroptosis.
Figures

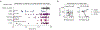
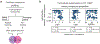

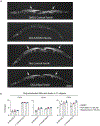
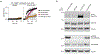



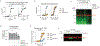






Comment in
-
STACKing the odds for discoveries.Nat Chem Biol. 2021 Jun;17(6):627-628. doi: 10.1038/s41589-021-00743-4. Nat Chem Biol. 2021. PMID: 33686293 No abstract available.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

