Molecular mapping of interstitial lung disease reveals a phenotypically distinct senescent basal epithelial cell population
- PMID: 33705361
- PMCID: PMC8119199
- DOI: 10.1172/jci.insight.143626
Molecular mapping of interstitial lung disease reveals a phenotypically distinct senescent basal epithelial cell population
Abstract
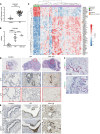
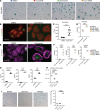
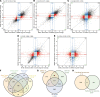
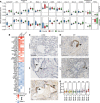
Compromised regenerative capacity of lung epithelial cells can lead to cellular senescence, which may precipitate fibrosis. While increased markers of senescence have been reported in idiopathic pulmonary fibrosis (IPF), the origin and identity of these senescent cells remain unclear, and tools to characterize context-specific cellular senescence in human lung are lacking. We observed that the senescent marker p16 is predominantly localized to bronchiolized epithelial structures in scarred regions of IPF and systemic sclerosis-associated interstitial lung disease (SSc-ILD) lung tissue, overlapping with the basal epithelial markers Keratin 5 and Keratin 17. Using in vitro models, we derived transcriptional signatures of senescence programming specific to different types of lung epithelial cells and interrogated these signatures in a single-cell RNA-Seq data set derived from control, IPF, and SSc-ILD lung tissue. We identified a population of basal epithelial cells defined by, and enriched for, markers of cellular senescence and identified candidate markers specific to senescent basal epithelial cells in ILD that can enable future functional studies. Notably, gene expression of these cells significantly overlaps with terminally differentiating cells in stratified epithelia, where it is driven by p53 activation as part of the senescence program.
Keywords: Aging; Cellular senescence; Fibrosis; Pulmonology.
Conflict of interest statement
Figures



References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

