Principles and practice of determining metal-protein affinities
- PMID: 33710331
- PMCID: PMC7959690
- DOI: 10.1042/BCJ20200838
Principles and practice of determining metal-protein affinities
Abstract
Metal ions play many critical roles in biology, as structural and catalytic cofactors, and as cell regulatory and signalling elements. The metal-protein affinity, expressed conveniently by the metal dissociation constant, KD, describes the thermodynamic strength of a metal-protein interaction and is a key parameter that can be used, for example, to understand how proteins may acquire metals in a cell and to identify dynamic elements (e.g. cofactor binding, changing metal availabilities) which regulate protein metalation in vivo. Here, we outline the fundamental principles and practical considerations that are key to the reliable quantification of metal-protein affinities. We review a selection of spectroscopic probes which can be used to determine protein affinities for essential biological transition metals (including Mn(II), Fe(II), Co(II), Ni(II), Cu(I), Cu(II) and Zn(II)) and, using selected examples, demonstrate how rational probe selection combined with prudent experimental design can be applied to determine accurate KD values.
Keywords: dissociation constant; metal probe; metalloprotein; metal–protein affinity; spectroscopic probe; transition metals.
© 2021 The Author(s).
Conflict of interest statement
The authors declare that there are no competing interests associated with the manuscript.
Figures

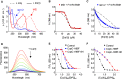
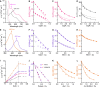



References
-
- Xiao, Z. and Wedd, A.G. (2021) Coping with toxic metals. Met. Ions Life Sci. 21, 271–298 10.3390/ijms161226183 - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

