Chemotherapy-Enriched THBS2-Deficient Cancer Stem Cells Drive Hepatocarcinogenesis through Matrix Softness Induced Histone H3 Modifications
- PMID: 33717837
- PMCID: PMC7927606
- DOI: 10.1002/advs.202002483
Chemotherapy-Enriched THBS2-Deficient Cancer Stem Cells Drive Hepatocarcinogenesis through Matrix Softness Induced Histone H3 Modifications
Abstract
The physical microenvironment is a critical mediator of tumor behavior. However, detailed biological and mechanistic insight is lacking. The present study reveals the role of chemotherapy-enriched CD133+ liver cancer stem cells (CSCs) with THBS2 deficiency. This subpopulation of cells contributes to a more aggressive cancer and functional stemness phenotype in hepatocellular carcinoma (HCC) by remodeling the extracellular matrix (ECM) through the regulation of matrix metalloproteinase (MMP) activity, collagen degradation, and matrix stiffness. The local soft spots created by these liver CSCs can enhance stemness and drug resistance and provide a route of escape to facilitate HCC metastasis. Interestingly, a positive feed-forward loop is identified where a local soft spot microenvironment in the HCC tumor is enriched with CD133 expressing cells that secrete markedly less ECM-modifying THBS2 upon histone H3 modification at its promoter region, allowing the maintenance of a localized soft spot matrix. Clinically, THBS2 deficiency is also correlated with low HCC survival, where high levels of CSCs with low THBS2 expression in HCC are associated with decreased collagen fiber deposits and an invasive tumor front. The findings have implications for the treatment of cancer stemness and for the prevention of tumor outgrowth through disseminated tumor cells.
Keywords: CD133; THBS2; cancer stemness; hepatocellular carcinomas; histone modifications; matrix stiffness; mechanoepigenetics; metastasis.
© 2021 The Authors. Advanced Science published by Wiley‐VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



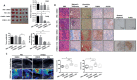


References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous
