Quantitative Proteomic Profiling Identifies a Potential Novel Chaperone Marker in Resistant Breast Cancer
- PMID: 33718123
- PMCID: PMC7951058
- DOI: 10.3389/fonc.2021.540134
Quantitative Proteomic Profiling Identifies a Potential Novel Chaperone Marker in Resistant Breast Cancer
Abstract
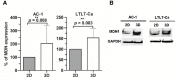
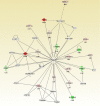
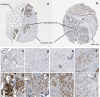
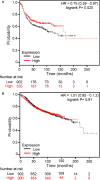
Development of aromatase inhibitor resistant breast cancer among postmenopausal women continues to be a major clinical obstacle. Previously, our group demonstrated that as breast cancer cells transition from hormone-dependent to hormone-independent, they are associated with increased growth factor signaling, enhanced cellular motility, and the epithelial to mesenchymal transition (EMT). Given the complexity of cancer stem cells (CSC) and their implications on endocrine resistance and EMT, we sought to understand their contribution towards the development of aromatase inhibitor resistant breast cancer. Cells cultured three dimensionally as mammospheres are enriched for CSCs and more accurately recapitulates tumors in vivo. Therefore, a global proteomic analysis was conducted using letrozole resistant breast cancer cells (LTLT-Ca) mammospheres and compared to their adherent counterparts. Results demonstrated over 1000 proteins with quantitative abundance ratios were identified. Among the quantified proteins, 359 were significantly altered (p < 0.05), where 173 were upregulated and 186 downregulated (p < 0.05, fold change >1.20). Notably, midasin, a chaperone protein required for maturation and nuclear export of the pre-60S ribosome was increased 35-fold. Protein expression analyses confirmed midasin is ubiquitously expressed in normal tissue but is overexpressed in lobular and ductal breast carcinoma tissue as well as ER+ and ER- breast cancer cell lines. Functional enrichment analyses indicated that 19 gene ontology terms and one KEGG pathway were over-represented by the down-regulated proteins and both were associated with protein synthesis. Increased midasin was strongly correlated with decreased relapse free survival in hormone independent breast cancer. For the first time, we characterized the global proteomic signature of CSC-enriched letrozole-resistant cells associated with protein synthesis, which may implicate a role for midasin in endocrine resistance.
Keywords: aromatase inhibitors; breast cancer; cancer stem cells; letrozole resistance; midasin; translation.
Copyright © 2021 Gallegos, Patel, Llopis, Walker, Davidson, Zhang, Zhang and Tilghman.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
Mammospheres of letrozole-resistant breast cancer cells enhance breast cancer aggressiveness.Oncol Lett. 2021 Aug;22(2):620. doi: 10.3892/ol.2021.12881. Epub 2021 Jun 28. Oncol Lett. 2021. PMID: 34267813 Free PMC article.
-
Proteomic signatures of acquired letrozole resistance in breast cancer: suppressed estrogen signaling and increased cell motility and invasiveness.Mol Cell Proteomics. 2013 Sep;12(9):2440-55. doi: 10.1074/mcp.M112.023861. Epub 2013 May 23. Mol Cell Proteomics. 2013. PMID: 23704778 Free PMC article.
-
Glyceollin I Reverses Epithelial to Mesenchymal Transition in Letrozole Resistant Breast Cancer through ZEB1.Int J Environ Res Public Health. 2015 Dec 22;13(1):ijerph13010010. doi: 10.3390/ijerph13010010. Int J Environ Res Public Health. 2015. PMID: 26703648 Free PMC article.
-
Anti-tumor effects of letrozole.Cancer Invest. 2002;20 Suppl 2:15-21. doi: 10.1081/cnv-120014882. Cancer Invest. 2002. PMID: 12442345 Review.
-
Structural and functional characterization of aromatase, estrogen receptor, and their genes in endocrine-responsive and -resistant breast cancer cells.J Steroid Biochem Mol Biol. 2016 Jul;161:73-83. doi: 10.1016/j.jsbmb.2015.07.018. Epub 2015 Aug 13. J Steroid Biochem Mol Biol. 2016. PMID: 26277097 Free PMC article. Review.
Cited by
-
A Novel Allosteric Inhibitor Targets PLK1 in Triple Negative Breast Cancer Cells.Biomolecules. 2022 Mar 31;12(4):531. doi: 10.3390/biom12040531. Biomolecules. 2022. PMID: 35454120 Free PMC article.
-
AEGAN-Pathifier: a data augmentation method to improve cancer classification for imbalanced gene expression data.BMC Bioinformatics. 2024 Dec 27;25(1):392. doi: 10.1186/s12859-024-06013-z. BMC Bioinformatics. 2024. PMID: 39731019 Free PMC article.
-
Highly conserved ribosome biogenesis pathways between human and yeast revealed by the MDN1-NLE1 interaction and NLE1 containing pre-60S subunits.Nucleic Acids Res. 2025 Apr 10;53(7):gkaf255. doi: 10.1093/nar/gkaf255. Nucleic Acids Res. 2025. PMID: 40207627 Free PMC article.
-
Acquisition of Letrozole Resistance Through Activation of the p38/MAPK Signaling Cascade.Anticancer Res. 2021 Feb;41(2):583-599. doi: 10.21873/anticanres.14810. Anticancer Res. 2021. PMID: 33517263 Free PMC article.
References
-
- Brodie A, Jelovac D, Macedo L, Sabnis G, Tilghman S, Goloubeva O. Therapeutic observations in MCF-7 aromatase xenografts. Clin Cancer Res (2005. a) 11(2 Pt 2):884s–8s. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

