Simple, Affordable, and Modular Patterning of Cells using DNA
- PMID: 33720126
- PMCID: PMC10870346
- DOI: 10.3791/61937
Simple, Affordable, and Modular Patterning of Cells using DNA
Abstract
The relative positioning of cells is a key feature of the microenvironment that organizes cell-cell interactions. To study the interactions between cells of the same or different type, micropatterning techniques have proved useful. DNA Programmed Assembly of Cells (DPAC) is a micropatterning technique that targets the adhesion of cells to a substrate or other cells using DNA hybridization. The most basic operations in DPAC begin with decorating cell membranes with lipid-modified oligonucleotides, then flowing them over a substrate that has been patterned with complementary DNA sequences. Cells adhere selectively to the substrate only where they find a complementary DNA sequence. Non-adherent cells are washed away, revealing a pattern of adherent cells. Additional operations include further rounds of cell-substrate or cell-cell adhesion, as well as transferring the patterns formed by DPAC to an embedding hydrogel for long-term culture. Previously, methods for patterning oligonucleotides on surfaces and decorating cells with DNA sequences required specialized equipment and custom DNA synthesis, respectively. We report an updated version of the protocol, utilizing an inexpensive benchtop photolithography setup and commercially available cholesterol modified oligonucleotides (CMOs) deployed using a modular format. CMO-labeled cells adhere with high efficiency to DNA-patterned substrates. This approach can be used to pattern multiple cell types at once with high precision and to create arrays of microtissues embedded within an extracellular matrix. Advantages of this method include its high resolution, ability to embed cells into a three-dimensional microenvironment without disrupting the micropattern, and flexibility in patterning any cell type.
Figures
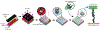


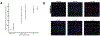

References
-
- Goubko C. a., Cao X Patterning multiple cell types in co-cultures: A review. Materials Science and Engineering C 29 (6), 1855–1868 (2009).
-
- Sun W et al. The bioprinting roadmap. Biofabrication 12 (2), 022002 (2020). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
