Stability Prediction for Mutations in the Cytosolic Domains of Cystic Fibrosis Transmembrane Conductance Regulator
- PMID: 33720715
- PMCID: PMC10230551
- DOI: 10.1021/acs.jcim.0c01207
Stability Prediction for Mutations in the Cytosolic Domains of Cystic Fibrosis Transmembrane Conductance Regulator
Abstract
Cystic Fibrosis (CF) is caused by mutations to the Cystic Fibrosis Transmembrane Conductance Regulator (CFTR) chloride channel. CFTR is composed of two membrane spanning domains, two cytosolic nucleotide-binding domains (NBD1 and NBD2) and a largely unstructured R-domain. Multiple CF-causing mutations reside in the NBDs and some are known to compromise the stability of these domains. The ability to predict the effect of mutations on the stability of the cytosolic domains of CFTR and to shed light on the mechanisms by which they exert their effect is therefore important in CF research. With this in mind, we have predicted the effect on domain stability of 59 mutations in NBD1 and NBD2 using 15 different algorithms and evaluated their performances via comparison to experimental data using several metrics including the correct classification rate (CCR), and the squared Pearson correlation (R2) and Spearman's correlation (ρ) calculated between the experimental ΔTm values and the computationally predicted ΔΔG values. Overall, the best results were obtained with FoldX and Rosetta. For NBD1 (35 mutations), FoldX provided R2 and ρ values of 0.64 and -0.71, respectively, with an 86% correct classification rate (CCR). For NBD2 (24 mutations), FoldX R2, ρ, and CCR were 0.51, -0.73, and 75%, respectively. Application of the Rosetta high-resolution protocol (Rosetta_hrp) to NBD1 yielded R2, ρ, and CCR of 0.64, -0.75, and 69%, respectively, and for NBD2 yielded R2, ρ, and CCR of 0.29, -0.27, and 50%, respectively. The corresponding numbers for the Rosetta's low-resolution protocol (Rosetta_lrp) were R2 = 0.47, ρ = -0.69, and CCR = 69% for NBD1 and R2 = 0.27, ρ = -0.24, and CCR = 63% for NBD2. For NBD1, both algorithms suggest that destabilizing mutations suffer from destabilizing vdW clashes, whereas stabilizing mutations benefit from favorable H-bond interactions. Two triple consensus approaches based on FoldX, Rosetta_lpr, and Rosetta_hpr were attempted using either "majority-voting" or "all-voting". The all-voting consensus outperformed the individual predictors, albeit on a smaller data set. In summary, our results suggest that the effect of mutations on the stability of CFTR's NBDs could be largely predicted. Since NBDs are common to all ABC transporters, these results may find use in predicting the effect and mechanism of the action of multiple disease-causing mutations in other proteins.
Conflict of interest statement
CONFLICT OF INTEREST
The authors declare no conflict of interest and do not endorse any program.
Figures
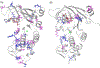
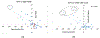
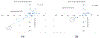

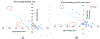

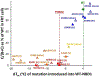
References
-
- Veit G; Avramescu RG; Chiang AN; Houck SA; Cai Z; Peters KW; Hong JS; Pollard HB; Guggino WB; Balch WE; Skach WR; Cutting GR; Frizzell RA; Sheppard DN; Cyr DM; Sorscher EJ; Brodsky JL; Lukacs GL From CFTR Biology toward Combinatorial Pharmacotherapy: Expanded Classification of Cystic Fibrosis Mutations. Mol. Biol. Cell 2016, 27, 424–433. 10.1091/mbc.e14-04-0935. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

