All-trans retinoic acid and protein kinase C α/β1 inhibitor combined treatment targets cancer stem cells and impairs breast tumor progression
- PMID: 33723318
- PMCID: PMC7961031
- DOI: 10.1038/s41598-021-85344-w
All-trans retinoic acid and protein kinase C α/β1 inhibitor combined treatment targets cancer stem cells and impairs breast tumor progression
Abstract
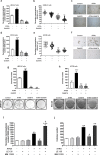
Breast cancer is the leading cause of cancer death among women worldwide. Blocking a single signaling pathway is often an ineffective therapy, especially in the case of aggressive or drug-resistant tumors. Since we have previously described the mechanism involved in the crosstalk between Retinoic Acid system and protein kinase C (PKC) pathway, the rationale of our study was to evaluate the effect of combining all-trans-retinoic acid (ATRA) with a classical PCK inhibitor (Gö6976) in preclinical settings. Employing hormone-independent mammary cancer models, Gö6976 and ATRA combined treatment induced a synergistic reduction in proliferative potential that correlated with an increased apoptosis and RARs modulation towards an anti-oncogenic profile. Combined treatment also impairs growth, self-renewal and clonogenicity potential of cancer stem cells and reduced tumor growth, metastatic spread and cancer stem cells frequency in vivo. An in-silico analysis of "Kaplan-Meier plotter" database indicated that low PKCα together with high RARα mRNA expression is a favorable prognosis factor for hormone-independent breast cancer patients. Here we demonstrate that a classical PKC inhibitor potentiates ATRA antitumor effects also targeting cancer stem cells growth, self-renewal and frequency.
Conflict of interest statement
The authors declare no competing interests.
Figures






Similar articles
-
Involvement of protein kinase C α and δ activities on the induction of the retinoic acid system in mammary cancer cells.Mol Carcinog. 2015 Oct;54(10):1110-21. doi: 10.1002/mc.22181. Epub 2014 May 17. Mol Carcinog. 2015. PMID: 24838400
-
Upregulation and activation of PKC alpha by ErbB2 through Src promotes breast cancer cell invasion that can be blocked by combined treatment with PKC alpha and Src inhibitors.Oncogene. 2006 Jun 1;25(23):3286-95. doi: 10.1038/sj.onc.1209361. Epub 2006 Jan 23. Oncogene. 2006. PMID: 16407820
-
PKCδ Inhibition Impairs Mammary Cancer Proliferative Capacity But Selects Cancer Stem Cells, Involving Autophagy.J Cell Biochem. 2016 Mar;117(3):730-40. doi: 10.1002/jcb.25358. Epub 2015 Sep 10. J Cell Biochem. 2016. PMID: 26335446
-
Myoepithelial and luminal breast cancer cells exhibit different responses to all-trans retinoic acid.Cell Oncol (Dordr). 2015 Aug;38(4):289-305. doi: 10.1007/s13402-015-0230-z. Epub 2015 Jun 5. Cell Oncol (Dordr). 2015. PMID: 26044847
-
Retinoic acid and the histone deacetylase inhibitor trichostatin a inhibit the proliferation of human renal cell carcinoma in a xenograft tumor model.Clin Cancer Res. 2005 May 1;11(9):3558-66. doi: 10.1158/1078-0432.CCR-04-1155. Clin Cancer Res. 2005. PMID: 15867260
Cited by
-
Targeting the retinoic acid signaling pathway as a modern precision therapy against cancers.Front Cell Dev Biol. 2023 Aug 11;11:1254612. doi: 10.3389/fcell.2023.1254612. eCollection 2023. Front Cell Dev Biol. 2023. PMID: 37645246 Free PMC article. Review.
-
Antineoplastic activity of products derived from cellulose-containing materials: levoglucosenone and structurally-related derivatives as new alternatives for breast cancer treatment.Invest New Drugs. 2022 Feb;40(1):30-41. doi: 10.1007/s10637-021-01167-6. Epub 2021 Sep 3. Invest New Drugs. 2022. PMID: 34478029
-
Combination Treatment of Retinoic Acid Plus Focal Adhesion Kinase Inhibitor Prevents Tumor Growth and Breast Cancer Cell Metastasis.Cells. 2022 Sep 26;11(19):2988. doi: 10.3390/cells11192988. Cells. 2022. PMID: 36230951 Free PMC article.
-
Microbiota-Derived Natural Products Targeting Cancer Stem Cells: Inside the Gut Pharma Factory.Int J Mol Sci. 2023 Mar 5;24(5):4997. doi: 10.3390/ijms24054997. Int J Mol Sci. 2023. PMID: 36902427 Free PMC article. Review.
-
Protein Kinase C (PKC) Isozymes as Diagnostic and Prognostic Biomarkers and Therapeutic Targets for Cancer.Cancers (Basel). 2022 Nov 3;14(21):5425. doi: 10.3390/cancers14215425. Cancers (Basel). 2022. PMID: 36358843 Free PMC article. Review.
References
-
- Ono M, Kosaka N, Tominaga N, Yoshioka Y, Takeshita F, Takahashi RU, Yoshida M, Tsuda H, Tamura K, Ochiya T. Exosomes from bone marrow mesenchymal stem cells contain a microRNA that promotes dormancy in metastatic breast cancer cells. Sci. Signal. 2014;7:ra63. doi: 10.1126/scisignal.2005231. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

