Detection of horizontal gene transfer in the genome of the choanoflagellate Salpingoeca rosetta
- PMID: 33727612
- PMCID: PMC7971027
- DOI: 10.1038/s41598-021-85259-6
Detection of horizontal gene transfer in the genome of the choanoflagellate Salpingoeca rosetta
Abstract
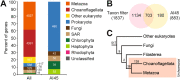
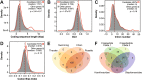
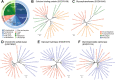
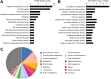
Horizontal gene transfer (HGT), the movement of heritable materials between distantly related organisms, is crucial in eukaryotic evolution. However, the scale of HGT in choanoflagellates, the closest unicellular relatives of metazoans, and its possible roles in the evolution of animal multicellularity remains unexplored. We identified at least 175 candidate HGTs in the genome of the colonial choanoflagellate Salpingoeca rosetta using sequence-based tests. The majority of these were orthologous to genes in bacterial and microalgal lineages, yet displayed genomic features consistent with the rest of the S. rosetta genome-evidence of ancient acquisition events. Putative functions include enzymes involved in amino acid and carbohydrate metabolism, cell signaling, and the synthesis of extracellular matrix components. Functions of candidate HGTs may have contributed to the ability of choanoflagellates to assimilate novel metabolites, thereby supporting adaptation, survival in diverse ecological niches, and response to external cues that are possibly critical in the evolution of multicellularity in choanoflagellates.
Conflict of interest statement
The authors declare no competing interests.
Figures
Similar articles
-
Choanoflagellate models - Monosiga brevicollis and Salpingoeca rosetta.Curr Opin Genet Dev. 2016 Aug;39:42-47. doi: 10.1016/j.gde.2016.05.016. Epub 2016 Jun 17. Curr Opin Genet Dev. 2016. PMID: 27318693 Review.
-
Genome editing enables reverse genetics of multicellular development in the choanoflagellate Salpingoeca rosetta.Elife. 2020 Jun 4;9:e56193. doi: 10.7554/eLife.56193. Elife. 2020. PMID: 32496191 Free PMC article.
-
The scale and evolutionary significance of horizontal gene transfer in the choanoflagellate Monosiga brevicollis.BMC Genomics. 2013 Oct 25;14(1):729. doi: 10.1186/1471-2164-14-729. BMC Genomics. 2013. PMID: 24156600 Free PMC article.
-
Premetazoan genome evolution and the regulation of cell differentiation in the choanoflagellate Salpingoeca rosetta.Genome Biol. 2013 Feb 18;14(2):R15. doi: 10.1186/gb-2013-14-2-r15. Genome Biol. 2013. PMID: 23419129 Free PMC article.
-
Horizontal gene transfer in choanoflagellates.J Exp Zool B Mol Dev Evol. 2013 Jan;320(1):1-9. doi: 10.1002/jez.b.22480. Epub 2012 Sep 19. J Exp Zool B Mol Dev Evol. 2013. PMID: 22997182 Review.
Cited by
-
Cross-phyla protein annotation by structural prediction and alignment.Genome Biol. 2023 May 12;24(1):113. doi: 10.1186/s13059-023-02942-9. Genome Biol. 2023. PMID: 37173746 Free PMC article.
-
Genomic insights into the evolution and adaptation of secondary metabolite gene clusters in fungicolous species Cladobotryum mycophilum ATHUM6906.G3 (Bethesda). 2024 Apr 3;14(4):jkae006. doi: 10.1093/g3journal/jkae006. G3 (Bethesda). 2024. PMID: 38214578 Free PMC article.
-
Metagenomic Insight into the Associated Microbiome in Plasmodia of Myxomycetes.Microorganisms. 2024 Dec 10;12(12):2540. doi: 10.3390/microorganisms12122540. Microorganisms. 2024. PMID: 39770743 Free PMC article.
-
Invasive mussels fashion silk-like byssus via mechanical processing of massive horizontally acquired coiled coils.Proc Natl Acad Sci U S A. 2023 Nov 28;120(48):e2311901120. doi: 10.1073/pnas.2311901120. Epub 2023 Nov 20. Proc Natl Acad Sci U S A. 2023. PMID: 37983489 Free PMC article.
-
A broad survey of choanoflagellates revises the evolutionary history of the Shaker family of voltage-gated K+ channels in animals.Proc Natl Acad Sci U S A. 2024 Jul 23;121(30):e2407461121. doi: 10.1073/pnas.2407461121. Epub 2024 Jul 17. Proc Natl Acad Sci U S A. 2024. PMID: 39018191 Free PMC article.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

