Single-cell analysis can define distinct evolution of tumor sites in follicular lymphoma
- PMID: 33728464
- PMCID: PMC8160505
- DOI: 10.1182/blood.2020009855
Single-cell analysis can define distinct evolution of tumor sites in follicular lymphoma
Abstract
Tumor heterogeneity complicates biomarker development and fosters drug resistance in solid malignancies. In lymphoma, our knowledge of site-to-site heterogeneity and its clinical implications is still limited. Here, we profiled 2 nodal, synchronously acquired tumor samples from 10 patients with follicular lymphoma (FL) using single-cell RNA, B-cell receptor (BCR) and T-cell receptor sequencing, and flow cytometry. By following the rapidly mutating tumor immunoglobulin genes, we discovered that BCR subclones were shared between the 2 tumor sites in some patients, but in many patients, the disease had evolved separately with limited tumor cell migration between the sites. Patients exhibiting divergent BCR evolution also exhibited divergent tumor gene-expression and cell-surface protein profiles. While the overall composition of the tumor microenvironment did not differ significantly between sites, we did detect a specific correlation between site-to-site tumor heterogeneity and T follicular helper (Tfh) cell abundance. We further observed enrichment of particular ligand-receptor pairs between tumor and Tfh cells, including CD40 and CD40LG, and a significant correlation between tumor CD40 expression and Tfh proliferation. Our study may explain discordant responses to systemic therapies, underscores the difficulty of capturing a patient's disease with a single biopsy, and furthers our understanding of tumor-immune networks in FL.
© 2021 by The American Society of Hematology.
Conflict of interest statement
Conflict-of-interest disclosure: The authors declare no competing financial interests.
Figures





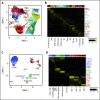


Comment in
-
Discordant solutions to discordant problems.Blood. 2021 May 27;137(21):2857-2858. doi: 10.1182/blood.2021011362. Blood. 2021. PMID: 34042983 No abstract available.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

