RNA-seq of human T cells after hematopoietic stem cell transplantation identifies Linc00402 as a regulator of T cell alloimmunity
- PMID: 33731431
- PMCID: PMC8589011
- DOI: 10.1126/scitranslmed.aaz0316
RNA-seq of human T cells after hematopoietic stem cell transplantation identifies Linc00402 as a regulator of T cell alloimmunity
Abstract
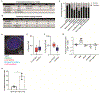
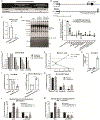
Mechanisms governing allogeneic T cell responses after solid organ and allogeneic hematopoietic stem cell transplantation (HSCT) are incompletely understood. To identify lncRNAs that regulate human donor T cells after clinical HSCT, we performed RNA sequencing on T cells from healthy individuals and donor T cells from three different groups of HSCT recipients that differed in their degree of major histocompatibility complex (MHC) mismatch. We found that lncRNA differential expression was greatest in T cells after MHC-mismatched HSCT relative to T cells after either MHC-matched or autologous HSCT. Differential expression was validated in an independent patient cohort and in mixed lymphocyte reactions using ex vivo healthy human T cells. We identified Linc00402, an uncharacterized lncRNA, among the lncRNAs differentially expressed between the mismatched unrelated and matched unrelated donor T cells. We found that Linc00402 was conserved and exhibited an 88-fold increase in human T cells relative to all other samples in the FANTOM5 database. Linc00402 was also increased in donor T cells from patients who underwent allogeneic cardiac transplantation and in murine T cells. Linc00402 was reduced in patients who subsequently developed acute graft-versus-host disease. Linc00402 enhanced the activity of ERK1 and ERK2, increased FOS nuclear accumulation, and augmented expression of interleukin-2 and Egr-1 after T cell receptor engagement. Functionally, Linc00402 augmented the T cell proliferative response to an allogeneic stimulus but not to a nominal ovalbumin peptide antigen or polyclonal anti-CD3/CD28 stimulus. Thus, our studies identified Linc00402 as a regulator of allogeneic T cell function.
Copyright © 2021 The Authors, some rights reserved; exclusive licensee American Association for the Advancement of Science. No claim to original U.S. Government Works.
Conflict of interest statement
Competing Interests
A.R. is member of Voxel Analytics, LLC. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




References
Publication types
MeSH terms
Substances
Grants and funding
- P30 CA046592/CA/NCI NIH HHS/United States
- R01 CA203542/CA/NCI NIH HHS/United States
- K12 HD028820/HD/NICHD NIH HHS/United States
- R01 CA217156/CA/NCI NIH HHS/United States
- R37 CA214955/CA/NCI NIH HHS/United States
- R01 AG028082/AG/NIA NIH HHS/United States
- T32 HL007622/HL/NHLBI NIH HHS/United States
- R01 AI138347/AI/NIAID NIH HHS/United States
- P30 AG024824/AG/NIA NIH HHS/United States
- T32 HG000040/HG/NHGRI NIH HHS/United States
- L40 HL138824/HL/NHLBI NIH HHS/United States
- T32 GM007863/GM/NIGMS NIH HHS/United States
- R01 HL128046/HL/NHLBI NIH HHS/United States
- UL1 TR002240/TR/NCATS NIH HHS/United States
- T32 AG062403/AG/NIA NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

