Genome-wide methylome analysis of two strains belonging to the hypervirulent Neisseria meningitidis serogroup W ST-11 clonal complex
- PMID: 33737546
- PMCID: PMC7973814
- DOI: 10.1038/s41598-021-85266-7
Genome-wide methylome analysis of two strains belonging to the hypervirulent Neisseria meningitidis serogroup W ST-11 clonal complex
Abstract
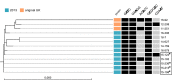
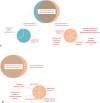
A rising incidence of meningococcal serogroup W disease has been evident in many countries worldwide. Serogroup W isolates belonging to the sequence type (ST)-11 clonal complex have been associated with atypical symptoms and increased case fatality rates. The continued expansion of this clonal complex in the later part of the 2010s has been largely due to a shift from the so-called original UK strain to the 2013 strain. Here we used single-molecule real-time (SMRT) sequencing to determine the methylomes of the two major serogroup W strains belonging to ST-11 clonal complex. Five methylated motifs were identified in this study, and three of the motifs, namely 5'-GATC-3', 5'-GAAGG-3', 5'-GCGCGC-3', were found in all 13 isolates investigated. The results showed no strain-specific motifs or difference in active restriction modification systems between the two strains. Two phase variable methylases were identified and the enrichment or depletion of the methylation motifs generated by these methylases varied between the two strains. Results from this work give further insight into the low diversity of methylomes in highly related strains and encourage further research to decipher the role of regions with under- or overrepresented methylation motifs.
Conflict of interest statement
RJR, BPA, and AF work for New England Biolabs, a company that sells research reagents including restriction enzymes and DNA methyltransferases to the scientific community. The commercial affiliation of these authors does not alter our adherence policies on sharing data and materials. All other authors have no conflicts to declare.
Figures



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

