Rubisco Adaptation Is More Limited by Phylogenetic Constraint Than by Catalytic Trade-off
- PMID: 33739416
- PMCID: PMC8233502
- DOI: 10.1093/molbev/msab079
Rubisco Adaptation Is More Limited by Phylogenetic Constraint Than by Catalytic Trade-off
Abstract
Rubisco assimilates CO2 to form the sugars that fuel life on earth. Correlations between rubisco kinetic traits across species have led to the proposition that rubisco adaptation is highly constrained by catalytic trade-offs. However, these analyses did not consider the phylogenetic context of the enzymes that were analyzed. Thus, it is possible that the correlations observed were an artefact of the presence of phylogenetic signal in rubisco kinetics and the phylogenetic relationship between the species that were sampled. Here, we conducted a phylogenetically resolved analysis of rubisco kinetics and show that there is a significant phylogenetic signal in rubisco kinetic traits. We re-evaluated the extent of catalytic trade-offs accounting for this phylogenetic signal and found that all were attenuated. Following phylogenetic correction, the largest catalytic trade-offs were observed between the Michaelis constant for CO2 and carboxylase turnover (∼21-37%), and between the Michaelis constants for CO2 and O2 (∼9-19%), respectively. All other catalytic trade-offs were substantially attenuated such that they were marginal (<9%) or non-significant. This phylogenetically resolved analysis of rubisco kinetic evolution also identified kinetic changes that occur concomitant with the evolution of C4 photosynthesis. Finally, we show that phylogenetic constraints have played a larger role than catalytic trade-offs in limiting the evolution of rubisco kinetics. Thus, although there is strong evidence for some catalytic trade-offs, rubisco adaptation has been more limited by phylogenetic constraint than by the combined action of all catalytic trade-offs.
Keywords: C4 photosynthesis; catalytic constraint; evolution; phylogenetic constraint; rubisco.
© The Author(s) 2021. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution.
Figures
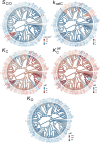
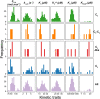



References
-
- Abouheif E. 1999. A method for testing the assumption of phylogenetic independence in comparative data. Evol Ecol Res. 1(8):895–909.
-
- Ackerly DD. 2004. Adaptation, niche conservatism, and convergence: comparative studies of leaf evolution in the California chaparral. Am Nat. 163(5):654–671. - PubMed
-
- Aigner H, Wilson RH, Bracher A, Calisse L, Bhat JY, Hartl FU, Hayer-Hartl M.. 2017. Plant RuBisCo assembly in E. coli with five chloroplast chaperones including BSD2’. Science 358(6368):1272–1278. - PubMed
-
- Andersson I, Backlund A.. 2008. Structure and function of Rubisco. Plant Physiol Biochem. 46(3):275–291. - PubMed
-
- Andrews TJ. 1988. Catalysis by cyanobacterial ribulose-bisphosphate carboxylase large subunits in the complete absence of small subunits. J Biol Chem. 263(25):12213–12219. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

