Biofilm Formation by Listeria monocytogenes 15G01, a Persistent Isolate from a Seafood-Processing Plant, Is Influenced by Inactivation of Multiple Genes Belonging to Different Functional Groups
- PMID: 33741610
- PMCID: PMC8117777
- DOI: 10.1128/AEM.02349-20
Biofilm Formation by Listeria monocytogenes 15G01, a Persistent Isolate from a Seafood-Processing Plant, Is Influenced by Inactivation of Multiple Genes Belonging to Different Functional Groups
Abstract
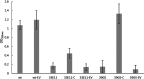
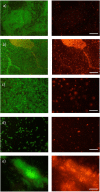
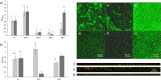
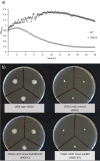
Listeria monocytogenes is a ubiquitous foodborne pathogen that results in a high rate of mortality in sensitive and immunocompromised people. Contamination of food with L. monocytogenes is thought to occur during food processing, most often as a result of the pathogen producing a biofilm that persists in the environment and acting as the source for subsequent dispersal of cells onto food. A survey of seafood-processing plants in New Zealand identified the persistent strain 15G01, which has a high capacity to form biofilms. In this study, a transposon library of L. monocytogenes 15G01 was screened for mutants with altered biofilm formation, assessed by a crystal violet assay, to identify genes involved in biofilm formation. This screen identified 36 transposants that showed a significant change in biofilm formation compared to the wild type. The insertion sites were in 27 genes, 20 of which led to decreased biofilm formation and seven to an increase. Two insertions were in intergenic regions. Annotation of the genes suggested that they are involved in diverse cellular processes, including stress response, autolysis, transporter systems, and cell wall/membrane synthesis. Analysis of the biofilms produced by the transposants using scanning electron microscopy and fluorescence microscopy showed notable differences in the structure of the biofilms compared to the wild type. In particular, inactivation of uvrB and mltD produced coccoid-shaped cells and elongated cells in long chains, respectively, and the mgtB mutant produced a unique biofilm with a sandwich structure which was reversed to the wild-type level upon magnesium addition. The mltD transposant was successfully complemented with the wild-type gene, whereas the phenotypes were not or only partially restored for the remaining mutants.IMPORTANCE The major source of contamination of food with Listeria monocytogenes is thought to be due to biofilm formation and/or persistence in food-processing plants. By establishing as a biofilm, L. monocytogenes cells become harder to eradicate due to their increased resistance to environmental threats. Understanding the genes involved in biofilm formation and their influence on biofilm structure will help identify new ways to eliminate harmful biofilms in food processing environments. To date, multiple genes have been identified as being involved in biofilm formation by L. monocytogenes; however, the exact mechanism remains unclear. This study identified four genes associated with biofilm formation by a persistent strain. Extensive microscopic analysis illustrated the effect of the disruption of mgtB, clsA, uvrB, and mltD and the influence of magnesium on the biofilm structure. The results strongly suggest an involvement in biofilm formation for the four genes and provide a basis for further studies to analyze gene regulation to assess the specific role of these biofilm-associated genes.
Keywords: Listeria monocytogenes; biofilms; food safety; mutagenesis.
Copyright © 2021 American Society for Microbiology.
Figures

Similar articles
-
Inactivation of the gene encoding the cationic antimicrobial peptide resistance factor MprF increases biofilm formation but reduces invasiveness of Listeria monocytogenes.J Appl Microbiol. 2021 Feb;130(2):464-477. doi: 10.1111/jam.14790. Epub 2020 Aug 10. J Appl Microbiol. 2021. PMID: 32687650
-
Genes involved in Listeria monocytogenes biofilm formation at a simulated food processing plant temperature of 15 °C.Int J Food Microbiol. 2016 Apr 16;223:63-74. doi: 10.1016/j.ijfoodmicro.2016.02.009. Epub 2016 Feb 11. Int J Food Microbiol. 2016. PMID: 26900648
-
Identification of Listeria monocytogenes determinants required for biofilm formation.PLoS One. 2014 Dec 17;9(12):e113696. doi: 10.1371/journal.pone.0113696. eCollection 2014. PLoS One. 2014. PMID: 25517120 Free PMC article.
-
Molecular biology of surface colonization by Listeria monocytogenes: an additional facet of an opportunistic Gram-positive foodborne pathogen.Environ Microbiol. 2011 Apr;13(4):835-50. doi: 10.1111/j.1462-2920.2010.02378.x. Epub 2010 Nov 18. Environ Microbiol. 2011. PMID: 21087384 Review.
-
Comprehensive strategies for controlling Listeria monocytogenes biofilms on food-contact surfaces.Compr Rev Food Sci Food Saf. 2024 May;23(3):e13348. doi: 10.1111/1541-4337.13348. Compr Rev Food Sci Food Saf. 2024. PMID: 38720587 Review.
Cited by
-
Nature-Inspired Antimicrobial Surfaces and Their Potential Applications in Food Industries.Foods. 2022 Mar 16;11(6):844. doi: 10.3390/foods11060844. Foods. 2022. PMID: 35327267 Free PMC article. Review.
-
Comparison of Selected Phenotypic Features of Persistent and Sporadic Strains of Listeria monocytogenes Sampled from Fish Processing Plants.Foods. 2022 May 20;11(10):1492. doi: 10.3390/foods11101492. Foods. 2022. PMID: 35627065 Free PMC article.
-
Persistence of microbiological hazards in food and feed production and processing environments.EFSA J. 2024 Jan 19;22(1):e8521. doi: 10.2903/j.efsa.2024.8521. eCollection 2024 Jan. EFSA J. 2024. PMID: 38250499 Free PMC article.
-
Catabolite control protein C contributes to virulence and hydrogen peroxide-induced oxidative stress responses in Listeria monocytogenes.Front Microbiol. 2024 May 31;15:1403694. doi: 10.3389/fmicb.2024.1403694. eCollection 2024. Front Microbiol. 2024. PMID: 38881664 Free PMC article.
References
-
- Fletcher G, Rogers M, Wong R. 1994. Survey of Listeria monocytogenes in New Zealand seafood. J Aquat Food Prod 3:13–24. 10.1300/J030v03n02_03. - DOI
-
- Srey S, Jahid IK, Ha S-D. 2013. Biofilm formation in food industries: a food safety concern. Food Control 31:572–585. 10.1016/j.foodcont.2012.12.001. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

