Filling in the Gaps in Metformin Biodegradation: a New Enzyme and a Metabolic Pathway for Guanylurea
- PMID: 33741630
- PMCID: PMC8208167
- DOI: 10.1128/AEM.03003-20
Filling in the Gaps in Metformin Biodegradation: a New Enzyme and a Metabolic Pathway for Guanylurea
Abstract
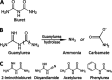
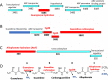
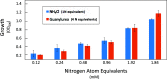
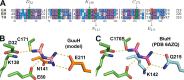
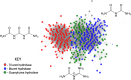
The widely prescribed pharmaceutical metformin and its main metabolite, guanylurea, are currently two of the most common contaminants in surface and wastewater. Guanylurea often accumulates and is poorly, if at all, biodegraded in wastewater treatment plants. This study describes Pseudomonas mendocina strain GU, isolated from a municipal wastewater treatment plant, using guanylurea as its sole nitrogen source. The genome was sequenced with 36-fold coverage and mined to identify guanylurea degradation genes. The gene encoding the enzyme initiating guanylurea metabolism was expressed, and the enzyme was purified and characterized. Guanylurea hydrolase, a newly described enzyme, was shown to transform guanylurea to one equivalent (each) of ammonia and guanidine. Guanidine also supports growth as a sole nitrogen source. Cell yields from growth on limiting concentrations of guanylurea revealed that metabolism releases all four nitrogen atoms. Genes encoding complete metabolic transformation were identified bioinformatically, defining the pathway as follows: guanylurea to guanidine to carboxyguanidine to allophanate to ammonia and carbon dioxide. The first enzyme, guanylurea hydrolase, is a member of the isochorismatase-like hydrolase protein family, which includes biuret hydrolase and triuret hydrolase. Although homologs, the three enzymes show distinct substrate specificities. Pairwise sequence comparisons and the use of sequence similarity networks allowed fine structure discrimination between the three homologous enzymes and provided insights into the evolutionary origins of guanylurea hydrolase.IMPORTANCE Metformin is a pharmaceutical most prescribed for type 2 diabetes and is now being examined for potential benefits to COVID-19 patients. People taking the drug pass it largely unchanged, and it subsequently enters wastewater treatment plants. Metformin has been known to be metabolized to guanylurea. The levels of guanylurea often exceed that of metformin, leading to the former being considered a "dead-end" metabolite. Metformin and guanylurea are water pollutants of emerging concern, as they persist to reach nontarget aquatic life and humans, the latter if it remains in treated water. The present study has identified a Pseudomonas mendocina strain that completely degrades guanylurea. The genome was sequenced, and the genes involved in guanylurea metabolism were identified in three widely separated genomic regions. This knowledge advances the idea that guanylurea is not a dead-end product and will allow for bioinformatic identification of the relevant genes in wastewater treatment plant microbiomes and other environments subjected to metagenomic sequencing.
Keywords: CgdAB; Pseudomonas mendocina; biodegradation; enzyme; genome; guanidine; guanylurea hydrolase; metformin.
Copyright © 2021 American Society for Microbiology.
Figures
Similar articles
-
Wastewater bacteria remediating the pharmaceutical metformin: Genomes, plasmids and products.Front Bioeng Biotechnol. 2022 Dec 16;10:1086261. doi: 10.3389/fbioe.2022.1086261. eCollection 2022. Front Bioeng Biotechnol. 2022. PMID: 36588930 Free PMC article.
-
Dinickel enzyme evolved to metabolize the pharmaceutical metformin and its implications for wastewater and human microbiomes.Proc Natl Acad Sci U S A. 2024 Mar 5;121(10):e2312652121. doi: 10.1073/pnas.2312652121. Epub 2024 Feb 26. Proc Natl Acad Sci U S A. 2024. PMID: 38408229 Free PMC article.
-
Solving the Conundrum: Widespread Proteins Annotated for Urea Metabolism in Bacteria Are Carboxyguanidine Deiminases Mediating Nitrogen Assimilation from Guanidine.Biochemistry. 2020 Sep 8;59(35):3258-3270. doi: 10.1021/acs.biochem.0c00537. Epub 2020 Aug 25. Biochemistry. 2020. PMID: 32786413 Free PMC article.
-
Co-occurrence, toxicity, and biotransformation pathways of metformin and its intermediate product guanylurea: Current state and future prospects for enhanced biodegradation strategy.Sci Total Environ. 2024 Apr 15;921:171108. doi: 10.1016/j.scitotenv.2024.171108. Epub 2024 Feb 22. Sci Total Environ. 2024. PMID: 38395159 Review.
-
Occurrence, Impact, Analysis and Treatment of Metformin and Guanylurea in Coastal Aquatic Environments of Canada, USA and Europe.Adv Mar Biol. 2018;81:23-58. doi: 10.1016/bs.amb.2018.09.005. Epub 2018 Nov 7. Adv Mar Biol. 2018. PMID: 30471658 Review.
Cited by
-
Growth of complete ammonia oxidizers on guanidine.Nature. 2024 Sep;633(8030):646-653. doi: 10.1038/s41586-024-07832-z. Epub 2024 Aug 14. Nature. 2024. PMID: 39143220 Free PMC article.
-
Discovery of a Ni2+-dependent heterohexameric metformin hydrolase.Nat Commun. 2024 Jul 20;15(1):6121. doi: 10.1038/s41467-024-50409-7. Nat Commun. 2024. PMID: 39033196 Free PMC article.
-
Combined effects of micropollutants and their degradation on prokaryotic communities at the sediment-water interface.Sci Rep. 2024 Jul 22;14(1):16840. doi: 10.1038/s41598-024-67308-y. Sci Rep. 2024. PMID: 39039186 Free PMC article.
-
Wastewater bacteria remediating the pharmaceutical metformin: Genomes, plasmids and products.Front Bioeng Biotechnol. 2022 Dec 16;10:1086261. doi: 10.3389/fbioe.2022.1086261. eCollection 2022. Front Bioeng Biotechnol. 2022. PMID: 36588930 Free PMC article.
-
Dinickel enzyme evolved to metabolize the pharmaceutical metformin and its implications for wastewater and human microbiomes.Proc Natl Acad Sci U S A. 2024 Mar 5;121(10):e2312652121. doi: 10.1073/pnas.2312652121. Epub 2024 Feb 26. Proc Natl Acad Sci U S A. 2024. PMID: 38408229 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

