Competition between bridged dinucleotides and activated mononucleotides determines the error frequency of nonenzymatic RNA primer extension
- PMID: 33744957
- PMCID: PMC8053118
- DOI: 10.1093/nar/gkab173
Competition between bridged dinucleotides and activated mononucleotides determines the error frequency of nonenzymatic RNA primer extension
Abstract
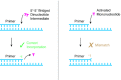
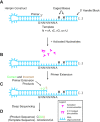
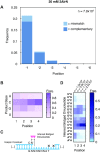
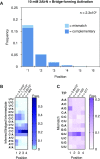
Nonenzymatic copying of RNA templates with activated nucleotides is a useful model for studying the emergence of heredity at the origin of life. Previous experiments with defined-sequence templates have pointed to the poor fidelity of primer extension as a major problem. Here we examine the origin of mismatches during primer extension on random templates in the simultaneous presence of all four 2-aminoimidazole-activated nucleotides. Using a deep sequencing approach that reports on millions of individual template-product pairs, we are able to examine correct and incorrect polymerization as a function of sequence context. We have previously shown that the predominant pathway for primer extension involves reaction with imidazolium-bridged dinucleotides, which form spontaneously by the reaction of two mononucleotides with each other. We now show that the sequences of correctly paired products reveal patterns that are expected from the bridged dinucleotide mechanism, whereas those associated with mismatches are consistent with direct reaction of the primer with activated mononucleotides. Increasing the ratio of bridged dinucleotides to activated mononucleotides, either by using purified components or by using isocyanide-based activation chemistry, reduces the error frequency. Our results point to testable strategies for the accurate nonenzymatic copying of arbitrary RNA sequences.
© The Author(s) 2021. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures


Similar articles
-
Enhanced nonenzymatic RNA copying with in-situ activation of short oligonucleotides.Nucleic Acids Res. 2023 Jul 21;51(13):6528-6539. doi: 10.1093/nar/gkad439. Nucleic Acids Res. 2023. PMID: 37247941 Free PMC article.
-
Insight into the mechanism of nonenzymatic RNA primer extension from the structure of an RNA-GpppG complex.Proc Natl Acad Sci U S A. 2017 Jul 18;114(29):7659-7664. doi: 10.1073/pnas.1704006114. Epub 2017 Jul 3. Proc Natl Acad Sci U S A. 2017. PMID: 28673998 Free PMC article.
-
Overcoming nucleotide bias in the nonenzymatic copying of RNA templates.Nucleic Acids Res. 2024 Dec 11;52(22):13515-13529. doi: 10.1093/nar/gkae982. Nucleic Acids Res. 2024. PMID: 39530216 Free PMC article.
-
The Mechanism of Nonenzymatic Template Copying with Imidazole-Activated Nucleotides.Angew Chem Int Ed Engl. 2019 Aug 5;58(32):10812-10819. doi: 10.1002/anie.201902050. Epub 2019 Jul 4. Angew Chem Int Ed Engl. 2019. PMID: 30908802 Review.
-
The Emergence of RNA from the Heterogeneous Products of Prebiotic Nucleotide Synthesis.J Am Chem Soc. 2021 Mar 10;143(9):3267-3279. doi: 10.1021/jacs.0c12955. Epub 2021 Feb 26. J Am Chem Soc. 2021. PMID: 33636080 Review.
Cited by
-
Diaminopurine in Nonenzymatic RNA Template Copying.J Am Chem Soc. 2024 Jun 12;146(23):15897-15907. doi: 10.1021/jacs.4c02560. Epub 2024 May 31. J Am Chem Soc. 2024. PMID: 38818863 Free PMC article.
-
Kinetic explanations for the sequence biases observed in the nonenzymatic copying of RNA templates.Nucleic Acids Res. 2022 Jan 11;50(1):35-45. doi: 10.1093/nar/gkab1202. Nucleic Acids Res. 2022. PMID: 34893864 Free PMC article.
-
Small-Molecule Organocatalysis Facilitates In Situ Nucleotide Activation and RNA Copying.J Am Chem Soc. 2023 Jul 26;145(29):16142-16149. doi: 10.1021/jacs.3c04635. Epub 2023 Jul 11. J Am Chem Soc. 2023. PMID: 37431761 Free PMC article.
-
Simulations predict preferred Mg2+ coordination in a nonenzymatic primer-extension reaction center.Biophys J. 2024 Jun 18;123(12):1579-1591. doi: 10.1016/j.bpj.2024.04.032. Epub 2024 May 3. Biophys J. 2024. PMID: 38702884 Free PMC article.
-
Unusual Base Pair between Two 2-Thiouridines and Its Implication for Nonenzymatic RNA Copying.J Am Chem Soc. 2024 Feb 14;146(6):3861-3871. doi: 10.1021/jacs.3c11158. Epub 2024 Jan 31. J Am Chem Soc. 2024. PMID: 38293747 Free PMC article.
References
-
- Gilbert W. Origin of life – the RNA world. Nature. 1986; 319:618–618.
-
- Orgel L.E. Was RNA the first genetic polymer?. Evol. Tinker. Gene Expr. 1989; 169:215–224.
-
- Joyce G.F. The antiquity of RNA-based evolution. Nature. 2002; 418:214–221. - PubMed
-
- Krishnamurthy R. On the emergence of RNA. Isr. J. Chem. 2015; 55:837–850.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

