Functional and Adaptive Significance of Promoter Mutations That Affect Divergent Myocardial Expressions of TRIM72 in Primates
- PMID: 33744959
- PMCID: PMC8233513
- DOI: 10.1093/molbev/msab083
Functional and Adaptive Significance of Promoter Mutations That Affect Divergent Myocardial Expressions of TRIM72 in Primates
Abstract
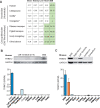
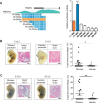
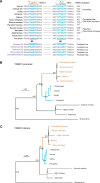
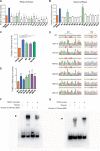
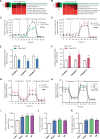
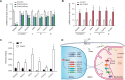
Cis-regulatory elements play important roles in tissue-specific gene expression and in the evolution of various phenotypes, and mutations in promoters and enhancers may be responsible for adaptations of species to environments. TRIM72 is a highly conserved protein that is involved in energy metabolism. Its expression in the heart varies considerably in primates, with high levels of expression in Old World monkeys and near absence in hominids. Here, we combine phylogenetic hypothesis testing and experimentation to demonstrate that mutations in promoter are responsible for the differences among primate species in the heart-specific expression of TRIM72. Maximum likelihood estimates of lineage-specific substitution rates under local-clock models show that relative to the evolutionary rate of introns, the rate of promoter was accelerated by 78% in the common ancestor of Old World monkeys, suggesting a role for positive selection in the evolution of the TRIM72 promoter, possibly driven by selective pressure due to changes in cardiac physiology after species divergence. We demonstrate that mutations in the TRIM72 promoter account for the differential myocardial TRIM72 expression of the human and the rhesus macaque. Furthermore, changes in TRIM72 expression alter the expression of genes involved in oxidative phosphorylation, which in turn affects mitochondrial respiration and cardiac energy capacity. On a broader timescale, phylogenetic regression analyses of data from 29 mammalian species show that mammals with high cardiac expression of TRIM72 have high heart rate, suggesting that the expression changes of TRIM72 may be related to differences in the heart physiology of those species.
Keywords: TRIM72; heart metabolism; primate evolution; promoter mutations.
© The Author(s) 2021. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution.
Figures
References
-
- Andolfatto P. 2005. Adaptive evolution of non-coding DNA in Drosophila. Nature 437(7062):1149–1152. - PubMed
-
- Brawand D, Soumillon M, Necsulea A, Julien P, Csardi G, Harrigan P, Weier M, Liechti A, Aximu-Petri A, Kircher M, et al. 2011. The evolution of gene expression levels in mammalian organs. Nature 478(7369):343–348. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

