Stochastic Simulation of Cellular Metabolism
- PMID: 33747671
- PMCID: PMC7971159
- DOI: 10.1109/access.2020.2986833
Stochastic Simulation of Cellular Metabolism
Abstract
Increased technological methods have enabled the investigation of biology at nanoscale levels. Such systems require the use of computational methods to comprehend the complex interactions that occur. The dynamics of metabolic systems have been traditionally described utilizing differential equations without fully capturing the heterogeneity of biological systems. Stochastic modeling approaches have recently emerged with the capacity to incorporate the statistical properties of such systems. However, the processing of stochastic algorithms is a computationally intensive task with intrinsic limitations. Alternatively, the queueing theory approach, historically used in the evaluation of telecommunication networks, can significantly reduce the computational power required to generate simulated results while simultaneously reducing the expansion of errors. We present here the application of queueing theory to simulate stochastic metabolic networks with high efficiency. With the use of glycolysis as a well understood biological model, we demonstrate the power of the proposed modeling methods discussed herein. Furthermore, we describe the simulation and pharmacological inhibition of glycolysis to provide an example of modeling capabilities.
Keywords: Biological Modeling; Glycolysis; Metabolic Networks; Metabolomics; Ordinary Differential Equations; Queueing Theory; Stochastic Simulation.
Figures




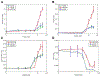

References
Grants and funding
LinkOut - more resources
Full Text Sources
