Simultaneous Ablation of the Catalytic AMPK α-Subunit SNF1 and Mitochondrial Matrix Protease CLPP Results in Pronounced Lifespan Extension
- PMID: 33748105
- PMCID: PMC7969656
- DOI: 10.3389/fcell.2021.616520
Simultaneous Ablation of the Catalytic AMPK α-Subunit SNF1 and Mitochondrial Matrix Protease CLPP Results in Pronounced Lifespan Extension
Abstract
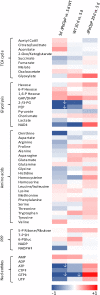
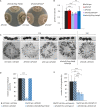
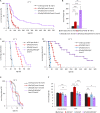
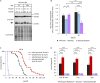
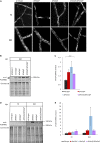
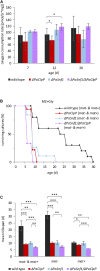
Organismic aging is known to be controlled by genetic and environmental traits. Pathways involved in the control of cellular metabolism play a crucial role. Previously, we identified a role of PaCLPP, a mitochondrial matrix protease, in the control of the mitochondrial energy metabolism, aging, and lifespan of the fungal aging model Podospora anserina. Most surprisingly, we made the counterintuitive observation that the ablation of this component of the mitochondrial quality control network leads to lifespan extension. In the current study, we investigated the role of energy metabolism of P. anserina. An age-dependent metabolome analysis of the wild type and a PaClpP deletion strain verified differences and changes of various metabolites in cultures of the PaClpP mutant and the wild type. Based on these data, we generated and analyzed a PaSnf1 deletion mutant and a ΔPaSnf1/ΔPaClpP double mutant. In both mutants PaSNF1, the catalytic α-subunit of AMP-activated protein kinase (AMPK) is ablated. PaSNF1 was found to be required for the development of fruiting bodies and ascospores and the progeny of sexual reproduction of this ascomycete and impact mitochondrial dynamics and autophagy. Most interestingly, while the single PaSnf1 deletion mutant is characterized by a slight lifespan increase, simultaneous deletion of PaSnf1 and PaClpP leads to a pronounced lifespan extension. This synergistic effect is strongly reinforced in the presence of the mating-type "minus"-linked allele of the rmp1 gene. Compared to the wild type, culture temperature of 35°C instead of the standard laboratory temperature of 27°C leads to a short-lived phenotype of the ΔPaSnf1/ΔPaClpP double mutant. Overall, our study provides novel evidence for complex interactions of different molecular pathways involved in mitochondrial quality control, gene expression, and energy metabolism in the control of organismic aging.
Keywords: AMPK; CLP protease; Podospora anserina; RMP1; SNF1; aging; development.
Copyright © 2021 Heinz, Krotova, Hamann and Osiewacz.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
Similar articles
-
Involvement of PaSNF1 in Fungal Development, Sterigmatocystin Biosynthesis, and Lignocellulosic Degradation in the Filamentous Fungus Podospora anserina.Front Microbiol. 2020 Jun 10;11:1038. doi: 10.3389/fmicb.2020.01038. eCollection 2020. Front Microbiol. 2020. PMID: 32587577 Free PMC article.
-
Disruption of the MICOS complex leads to an aberrant cristae structure and an unexpected, pronounced lifespan extension in Podospora anserina.J Cell Biochem. 2022 Aug;123(8):1306-1326. doi: 10.1002/jcb.30278. Epub 2022 May 26. J Cell Biochem. 2022. PMID: 35616269
-
Autophagy compensates impaired energy metabolism in CLPXP-deficient Podospora anserina strains and extends healthspan.Aging Cell. 2017 Aug;16(4):704-715. doi: 10.1111/acel.12600. Epub 2017 Apr 27. Aging Cell. 2017. PMID: 28449241 Free PMC article.
-
Mitochondrial metabolism and aging in the filamentous fungus Podospora anserina.Biochim Biophys Acta. 2006 May-Jun;1757(5-6):604-10. doi: 10.1016/j.bbabio.2006.03.005. Epub 2006 Mar 30. Biochim Biophys Acta. 2006. PMID: 16624249 Review.
-
Cellular copper homeostasis, mitochondrial DNA instabilities, and lifespan control in the filamentous fungus Podospora anserina.Exp Gerontol. 2000 Sep;35(6-7):677-86. doi: 10.1016/s0531-5565(00)00142-x. Exp Gerontol. 2000. PMID: 11053657 Review.
Cited by
-
Chemical tools to expand the ligandable proteome: Diversity-oriented synthesis-based photoreactive stereoprobes.Cell Chem Biol. 2024 Dec 19;31(12):2138-2155.e32. doi: 10.1016/j.chembiol.2024.10.005. Epub 2024 Nov 14. Cell Chem Biol. 2024. PMID: 39547236
-
A Network of Pathways Controlling Cellular Homeostasis Affects the Onset of Senescence in Podospora anserina.J Fungi (Basel). 2021 Mar 31;7(4):263. doi: 10.3390/jof7040263. J Fungi (Basel). 2021. PMID: 33807190 Free PMC article. Review.
-
Inactivity of Peptidase ClpP Causes Primary Accumulation of Mitochondrial Disaggregase ClpX with Its Interacting Nucleoid Proteins, and of mtDNA.Cells. 2021 Nov 29;10(12):3354. doi: 10.3390/cells10123354. Cells. 2021. PMID: 34943861 Free PMC article.
-
CLPP-Null Eukaryotes with Excess Heme Biosynthesis Show Reduced L-arginine Levels, Probably via CLPX-Mediated OAT Activation.Biomolecules. 2024 Feb 19;14(2):241. doi: 10.3390/biom14020241. Biomolecules. 2024. PMID: 38397478 Free PMC article.
-
Mitochondrial Phospholipid Homeostasis Is Regulated by the i-AAA Protease PaIAP and Affects Organismic Aging.Cells. 2021 Oct 16;10(10):2775. doi: 10.3390/cells10102775. Cells. 2021. PMID: 34685755 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

