Dual regulation of TxNIP by ChREBP and FoxO1 in liver
- PMID: 33748706
- PMCID: PMC7966993
- DOI: 10.1016/j.isci.2021.102218
Dual regulation of TxNIP by ChREBP and FoxO1 in liver
Abstract
TxNIP (Thioredoxin-interacting protein) is considered as a potential drug target for type 2 diabetes. Although TxNIP expression is correlated with hyperglycemia and glucotoxicity in pancreatic β cells, its regulation in liver cells has been less investigated. In the current study, we aim at providing a better understanding of Txnip regulation in hepatocytes in response to physiological stimuli and in the context of hyperglycemia in db/db mice. We focused on regulatory pathways governed by ChREBP (Carbohydrate Responsive Element Binding Protein) and FoxO1 (Forkhead box protein O1), transcription factors that play central roles in mediating the effects of glucose and fasting on gene expression, respectively. Studies using genetically modified mice reveal that hepatic TxNIP is up-regulated by both ChREBP and FoxO1 in liver cells and that its expression strongly correlates with fasting, suggesting a major role for this protein in the physiological adaptation to nutrient restriction.
Keywords: Endocrinology; Molecular Biology; Molecular Physiology.
© 2021.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





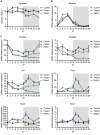


References
-
- Abdul-Wahed A., Guilmeau S., Postic C. Sweet sixteenth for ChREBP: established roles and future goals. Cell Metab. 2017;26:324–341. - PubMed
-
- Bedarida T., Baron S., Vibert F., Ayer A., Henrion D., Thioulouse E., Marchiol C., Beaudeux J.L., Cottart C.H., Nivet-Antoine V. Resveratrol decreases TXNIP mRNA and protein nuclear expressions with an arterial function improvement in old mice. J. Gerontol. A. Biol. Sci. Med. Sci. 2016;71:720–729. - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

