Protein Dynamics Influence the Enzymatic Activity of Phospholipase A/Acyltransferases 3 and 4
- PMID: 33749246
- PMCID: PMC8154263
- DOI: 10.1021/acs.biochem.0c00974
Protein Dynamics Influence the Enzymatic Activity of Phospholipase A/Acyltransferases 3 and 4
Abstract
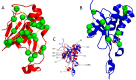
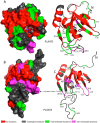
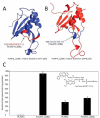
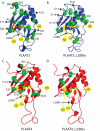
Phospholipase A/acyltransferase 3 (PLAAT3) and PLAAT4 are enzymes involved in the synthesis of bioactive lipids. Despite sequential and structural similarities, the two enzymes differ in activity and specificity. The relation between the activity and dynamics of the N-terminal domains of PLAAT3 and PLAAT4 was studied. PLAAT3 has a much higher melting temperature and exhibits less nanosecond and millisecond dynamics in the active site, in particular in loop L2(B6), as shown by NMR spectroscopy and molecular dynamics calculations. Swapping the L2(B6) loops between the two PLAAT enzymes results in strongly increased phospholipase activity in PLAAT3 but no reduction in PLAAT4 activity, indicating that this loop contributes to the low activity of PLAAT3. The results show that, despite structural similarity, protein dynamics differ substantially between the PLAAT variants, which can help to explain the activity and specificity differences.
Conflict of interest statement
The authors declare no competing financial interest.
Figures



References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

