Selective CRAF Inhibition Elicits Transactivation
- PMID: 33750116
- PMCID: PMC8041278
- DOI: 10.1021/jacs.0c11958
Selective CRAF Inhibition Elicits Transactivation
Abstract
Discovering molecules that regulate closely related protein isoforms is challenging, and in many cases the consequences of isoform-specific pharmacological regulation remains unknown. RAF isoforms are commonly mutated oncogenes that serve as effector kinases in MAP kinase signaling. BRAF/CRAF heterodimers are believed to be the primary RAF signaling species, and many RAF inhibitors lead to a "paradoxical activation" of RAF kinase activity through transactivation of the CRAF protomer; this leads to resistance mechanisms and secondary tumors. It has been hypothesized that CRAF-selective inhibition might bypass paradoxical activation, but no CRAF-selective inhibitor has been reported and the consequences of pharmacologically inhibiting CRAF have remained unknown. Here, we use bio-orthogonal ligand tethering (BOLT) to selectively target inhibitors to CRAF. Our results suggest that selective CRAF inhibition promotes paradoxical activation and exemplify how BOLT may be used to triage potential targets for drug discovery before any target-selective small molecules are known.
Conflict of interest statement
The authors declare the following competing financial interest(s): A.P.T., I.L.D., and J.H. are employees of AstraZeneca.
Figures


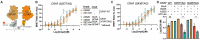
References
-
- Bishop A.; Buzko O.; Heyeck-Dumas S.; Jung I.; Kraybill B.; Liu Y.; Shah K.; Ulrich S.; Witucki L.; Yang F.; Zhang C.; Shokat K. M. Unnatural ligands for engineered proteins: new tools for chemical genetics. Annu. Rev. Biophys. Biomol. Struct. 2000, 29, 577–606. 10.1146/annurev.biophys.29.1.577. - DOI - PubMed
-
- Baud M. G. J.; Lin-Shiao E.; Cardote T.; Tallant C.; Pschibul A.; Chan K. H.; Zengerle M.; Garcia J. R.; Kwan T. T.; Ferguson F. M.; Ciulli A. Chemical biology. A bump-and-hole approach to engineer controlled selectivity of BET bromodomain chemical probes. Science (Washington, DC, U. S.) 2014, 346 (6209), 638–641. 10.1126/science.1249830. - DOI - PMC - PubMed
-
- Schumann B.; Malaker S. A.; Wisnovsky S. P.; Debets M. F.; Agbay A. J.; Fernandez D.; Wagner L. J. S.; Lin L.; Li Z.; Choi J.; Fox D. M.; Peh J.; Gray M. A.; Pedram K.; Kohler J. J.; Mrksich M.; Bertozzi C. R. Bump-and-Hole Engineering Identifies Specific Substrates of Glycosyltransferases in Living Cells. Mol. Cell 2020, 78 (5), 824–834.e15. 10.1016/j.molcel.2020.03.030. - DOI - PMC - PubMed
-
- Elliott T. S.; Townsley F. M.; Bianco A.; Ernst R. J.; Sachdeva A.; Elsasser S. J.; Davis L.; Lang K.; Pisa R.; Greiss S.; Lilley K. S.; Chin J. W. Proteome labeling and protein identification in specific tissues and at specific developmental stages in an animal. Nat. Biotechnol. 2014, 32 (5), 465–72. 10.1038/nbt.2860. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

