Transcriptomic analysis of feminizing somatic stem cells in the Drosophila testis reveals putative downstream effectors of the transcription factor Chinmo
- PMID: 33751104
- PMCID: PMC8759813
- DOI: 10.1093/g3journal/jkab067
Transcriptomic analysis of feminizing somatic stem cells in the Drosophila testis reveals putative downstream effectors of the transcription factor Chinmo
Abstract
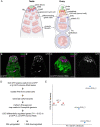
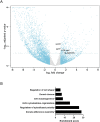
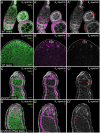
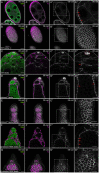
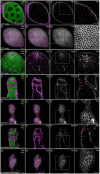
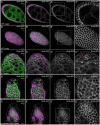
One of the best examples of sexual dimorphism is the development and function of the gonads, ovaries and testes, which produce sex-specific gametes, oocytes, and spermatids, respectively. The development of these specialized germ cells requires sex-matched somatic support cells. The sexual identity of somatic gonadal cells is specified during development and must be actively maintained during adulthood. We previously showed that the transcription factor Chinmo is required to ensure the male sexual identity of somatic support cells in the Drosophila melanogaster testis. Loss of chinmo from male somatic gonadal cells results in feminization: they transform from squamous to epithelial-like cells that resemble somatic cells in the female gonad but fail to properly ensheath the male germline, causing infertility. To identify potential target genes of Chinmo, we purified somatic cells deficient for chinmo from the adult Drosophila testis and performed next-generation sequencing to compare their transcriptome to that of control somatic cells. Bioinformatics revealed 304 and 1549 differentially upregulated and downregulated genes, respectively, upon loss of chinmo in early somatic cells. Using a combination of methods, we validated several differentially expressed genes. These data sets will be useful resources to the community.
Keywords: Chinmo; ovary; sex transformation; sexual identity; testis.
© The Author(s) 2021. Published by Oxford University Press on behalf of Genetics Society of America.
Figures

Similar articles
-
Chigno/CG11180 and SUMO are Chinmo-interacting proteins with a role in Drosophila testes somatic support cells.PeerJ. 2024 Mar 14;12:e16971. doi: 10.7717/peerj.16971. eCollection 2024. PeerJ. 2024. PMID: 38495765 Free PMC article.
-
Chinmo is sufficient to induce male fate in somatic cells of the adult Drosophila ovary.Development. 2016 Mar 1;143(5):754-63. doi: 10.1242/dev.129627. Epub 2016 Jan 25. Development. 2016. PMID: 26811385 Free PMC article.
-
The Jak-STAT target Chinmo prevents sex transformation of adult stem cells in the Drosophila testis niche.Dev Cell. 2014 Nov 24;31(4):474-86. doi: 10.1016/j.devcel.2014.10.004. Epub 2014 Nov 13. Dev Cell. 2014. PMID: 25453558 Free PMC article.
-
Genetic control of germline sexual dimorphism in Drosophila.Int Rev Cytol. 2002;219:1-60. doi: 10.1016/s0074-7696(02)19010-3. Int Rev Cytol. 2002. PMID: 12211627 Review.
-
Hedgehog in the Drosophila testis niche: what does it do there?Protein Cell. 2013 Sep;4(9):650-5. doi: 10.1007/s13238-013-3040-y. Epub 2013 Jun 26. Protein Cell. 2013. PMID: 23807635 Free PMC article. Review.
Cited by
-
Enrichment of Undifferentiated Germline and Somatic Cells from Drosophila Testes.Methods Mol Biol. 2023;2677:127-138. doi: 10.1007/978-1-0716-3259-8_7. Methods Mol Biol. 2023. PMID: 37464239
-
Soma-germline communication drives sex maintenance in the Drosophila testis.Natl Sci Rev. 2024 Jun 22;11(8):nwae215. doi: 10.1093/nsr/nwae215. eCollection 2024 Aug. Natl Sci Rev. 2024. PMID: 39183747 Free PMC article.
-
chinmo-mutant spermatogonial stem cells cause mitotic drive by evicting non-mutant neighbors from the niche.Dev Cell. 2022 Jan 10;57(1):80-94.e7. doi: 10.1016/j.devcel.2021.12.004. Epub 2021 Dec 22. Dev Cell. 2022. PMID: 34942115 Free PMC article.
-
Sex-converted testis somatic cells acquire female-specific behaviors and alter XY germline identity.Development. 2025 Aug 1;152(15):dev204785. doi: 10.1242/dev.204785. Epub 2025 Aug 7. Development. 2025. PMID: 40600835 Free PMC article.
-
Chigno/CG11180 and SUMO are Chinmo-interacting proteins with a role in Drosophila testes somatic support cells.PeerJ. 2024 Mar 14;12:e16971. doi: 10.7717/peerj.16971. eCollection 2024. PeerJ. 2024. PMID: 38495765 Free PMC article.
References
-
- Albertson R, Chabu C, Sheehan A, Doe CQ.. 2004. Scribble protein domain mapping reveals a multistep localization mechanism and domains necessary for establishing cortical polarity. J Cell Sci. 117:6061–6070. - PubMed
Publication types
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
