Ancestral reconstruction of mammalian FMO1 enables structural determination, revealing unique features that explain its catalytic properties
- PMID: 33759784
- PMCID: PMC7948450
- DOI: 10.1074/jbc.RA120.016297
Ancestral reconstruction of mammalian FMO1 enables structural determination, revealing unique features that explain its catalytic properties
Abstract
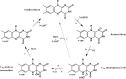
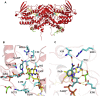
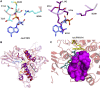
Mammals rely on the oxidative flavin-containing monooxygenases (FMOs) to detoxify numerous and potentially deleterious xenobiotics; this activity extends to many drugs, giving FMOs high pharmacological relevance. However, our knowledge regarding these membrane-bound enzymes has been greatly impeded by the lack of structural information. We anticipated that ancestral-sequence reconstruction could help us identify protein sequences that are more amenable to structural analysis. As such, we hereby reconstructed the mammalian ancestral protein sequences of both FMO1 and FMO4, denoted as ancestral flavin-containing monooxygenase (AncFMO)1 and AncFMO4, respectively. AncFMO1, sharing 89.5% sequence identity with human FMO1, was successfully expressed as a functional enzyme. It displayed typical FMO activities as demonstrated by oxygenating benzydamine, tamoxifen, and thioanisole, drug-related compounds known to be also accepted by human FMO1, and both NADH and NADPH cofactors could act as electron donors, a feature only described for the FMO1 paralogs. AncFMO1 crystallized as a dimer and was structurally resolved at 3.0 Å resolution. The structure harbors typical FMO aspects with the flavin adenine dinucleotide and NAD(P)H binding domains and a C-terminal transmembrane helix. Intriguingly, AncFMO1 also contains some unique features, including a significantly porous and exposed active site, and NADPH adopting a new conformation with the 2'-phosphate being pushed inside the NADP+ binding domain instead of being stretched out in the solvent. Overall, the ancestrally reconstructed mammalian AncFMO1 serves as the first structural model to corroborate and rationalize the catalytic properties of FMO1.
Keywords: NAD(P)H; ancestral-sequence reconstruction (ASR); crystal structure; enzyme kinetics; flavin adenine dinucleotide (FAD); flavin-containing monooxygenase (FMO); stopped flow.
Copyright © 2020 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Conflict of interest The authors declare that they have no conflicts of interest with the contents of this article.
Figures
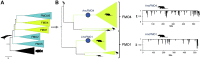
 ), bony fishes (
), bony fishes ( ), mammals (
), mammals ( ), aves (
), aves ( ) and testudines (
) and testudines ( ). Reconstructed mammalian ancestors are shown with blue circles. On the right the corresponding graphs of the posterior probability distribution per site are shown.
). Reconstructed mammalian ancestors are shown with blue circles. On the right the corresponding graphs of the posterior probability distribution per site are shown.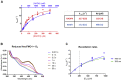
References
-
- Phillips I.R., Shephard E.A. Drug metabolism by flavin-containing monooxygenases of human and mouse. Expert Opin. Drug Metab. Toxicol. 2017;13:167–181. - PubMed
-
- Cruciani G., Valeri A., Goracci L., Pellegrino R.M., Buonerba F., Baroni M. Flavin monooxygenase metabolism: why medicinal chemists should matter. J. Med. Chem. 2014;57:6183–6196. - PubMed
-
- Shephard E.A., Dolphin C.T., Fox M.F., Povey S., Smith R., Phillips I.R. Localization of genes encoding three distinct flavin-containing monooxygenases to human chromosome 1q. Genomics. 1993;16:85–89. - PubMed
-
- McCombie R.R., Dolphin C.T., Povey S., Phillips I.R., Shephard E.A. Localization of human flavin-containing monooxygenase genes FMO2 and FMO5 to chromosome 1q. Genomics. 1996;34:426–429. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

