Homology Model of a Catalytically Competent Bifunctional Rel Protein
- PMID: 33763451
- PMCID: PMC7983052
- DOI: 10.3389/fmolb.2021.628596
Homology Model of a Catalytically Competent Bifunctional Rel Protein
Abstract
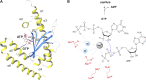
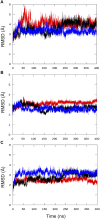
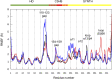
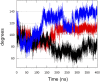
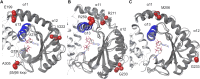
Bacteria have developed different bet hedging strategies to survive hostile environments and stressful conditions with persistency being maybe the most elegant yet still poorly understood one. Persisters' temporary tolerance to antibiotic treatment hints at their role not only in chronic and recurrent infections but also in the insurgence of resistant strains. Therefore, hampering persisters formation might represent an innovative strategy in the quest for new effective antimicrobial compounds. Among the molecular mechanisms postulated for the persister phenotypic switch, we decided to focus our attention on the stringent response and, in particular, on the upstream triggering step that is the accumulation of guanosine tetra- and pentaphosphate, collectivity called (p)ppGpp. Intracellular levels of (p)ppGpp are regulated by a superfamily of enzymes called RSH (RelA/SpoT homologue) that are able to promote its synthesis via pyrophosphate transfer from an ATP molecule to the 3' position of either GDP or GTP. These enzymes are classified based on the structural domain(s) present (only synthetase, only hydrolase, or both). Here we present our work on Rel Seq (from S. equisimilis), still the only bifunctional Rel protein for which a GDP-bound "synthetase-ON" structure is available. Analysis of the synthetase site, occupied only by GDP, revealed a partially active state, where the supposed ATP binding region is not conformationally apt to accommodate it. In order to achieve a protein model that gets closer to a fully active state, we generated a chimera structure of Rel Seq by homology modeling, starting from the crystal structure of the catalytically competent state of RelP, a smaller, single-domain, Rel protein from S. aureus. Molecular dynamics simulations allowed verifying the stability of the generated chimera model. Virtual screening and ligand design studies are underway.
Keywords: (p)ppGpp; RelP; RelSeq; bacterial persisters; chimera; homology modeling; molecular dynamics.
Copyright © 2021 Civera and Sattin.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
New Chemotypes for the Inhibition of (p)ppGpp Synthesis in the Quest for New Antimicrobial Compounds.Molecules. 2022 May 12;27(10):3097. doi: 10.3390/molecules27103097. Molecules. 2022. PMID: 35630574 Free PMC article.
-
Conformational antagonism between opposing active sites in a bifunctional RelA/SpoT homolog modulates (p)ppGpp metabolism during the stringent response [corrected].Cell. 2004 Apr 2;117(1):57-68. doi: 10.1016/s0092-8674(04)00260-0. Cell. 2004. PMID: 15066282
-
The Ps and Qs of alarmone synthesis in Staphylococcus aureus.PLoS One. 2019 Oct 15;14(10):e0213630. doi: 10.1371/journal.pone.0213630. eCollection 2019. PLoS One. 2019. PMID: 31613897 Free PMC article.
-
(p)ppGpp Metabolism and Antimicrobial Resistance in Bacterial Pathogens.Front Microbiol. 2020 Oct 9;11:563944. doi: 10.3389/fmicb.2020.563944. eCollection 2020. Front Microbiol. 2020. PMID: 33162948 Free PMC article. Review.
-
Stringent response protein as a potential target to intervene persistent bacterial infection.Biochimie. 2019 Oct;165:67-75. doi: 10.1016/j.biochi.2019.07.006. Epub 2019 Jul 11. Biochimie. 2019. PMID: 31302165 Review.
Cited by
-
New Chemotypes for the Inhibition of (p)ppGpp Synthesis in the Quest for New Antimicrobial Compounds.Molecules. 2022 May 12;27(10):3097. doi: 10.3390/molecules27103097. Molecules. 2022. PMID: 35630574 Free PMC article.
-
Many birds with one stone: targeting the (p)ppGpp signaling pathway of bacteria to improve antimicrobial therapy.Biophys Rev. 2021 Nov 12;13(6):1039-1051. doi: 10.1007/s12551-021-00895-6. eCollection 2021 Dec. Biophys Rev. 2021. PMID: 35059026 Free PMC article. Review.
References
-
- Berendsen H. J. C., Postma J. P. M., van Gunsteren W. F., DiNola A., Haak J. R. (1984). Molecular dynamics with coupling to an external bath. J. Chem. Phys. 81 (8), 3684–3690. 10.1063/1.448118 - DOI
-
- Case D. A. I. Y. B.-S., Brozell S. R., Cerutti D. S., Cheatham T. E., III, Cruzeiro V. W. D., et al. (2018). Amber 2018. San Francisco, CA: University of California.
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

