In silico investigation of critical binding pattern in SARS-CoV-2 spike protein with angiotensin-converting enzyme 2
- PMID: 33767306
- PMCID: PMC7994905
- DOI: 10.1038/s41598-021-86380-2
In silico investigation of critical binding pattern in SARS-CoV-2 spike protein with angiotensin-converting enzyme 2
Abstract
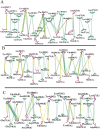
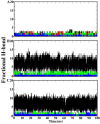
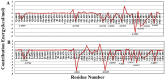
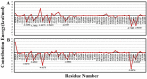
Severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) is a newly-discovered coronavirus and responsible for the spread of coronavirus disease 2019 (COVID-19). SARS-CoV-2 infected millions of people in the world and immediately became a pandemic in March 2020. SARS-CoV-2 belongs to the beta-coronavirus genus of the large family of Coronaviridae. It is now known that its surface spike glycoprotein binds to the angiotensin-converting enzyme-2 (ACE2), which is expressed on the lung epithelial cells, mediates the fusion of the cellular and viral membranes, and facilitates the entry of viral genome to the host cell. Therefore, blocking the virus-cell interaction could be a potential target for the prevention of viral infection. The binding of SARS-CoV-2 to ACE2 is a protein-protein interaction, and so, analyzing the structure of the spike glycoprotein of SARS-CoV-2 and its underlying mechanism to bind the host cell receptor would be useful for the management and treatment of COVID-19. In this study, we performed comparative in silico studies to deeply understand the structural and functional details of the interaction between the spike glycoprotein of SARS-CoV-2 and its cognate cellular receptor ACE2. According to our results, the affinity of the ACE2 receptor for SARS-CoV-2 was higher than SARS-CoV. According to the free energy decomposition of the spike glycoprotein-ACE2 complex, we found critical points in three areas which are responsible for the increased binding affinity of SARS-CoV-2 compared with SARS-CoV. These mutations occurred at the receptor-binding domain of the spike glycoprotein that play an essential role in the increasing the affinity of coronavirus to ACE2. For instance, mutations Pro462Ala and Leu472Phe resulted in the altered binding energy from - 2 kcal mol-1 in SARS-COV to - 6 kcal mol-1 in SARS-COV-2. The results demonstrated that some mutations in the receptor-binding motif could be considered as a hot-point for designing potential drugs to inhibit the interaction between the spike glycoprotein and ACE2.
Conflict of interest statement
The authors declare no competing interests.
Figures

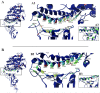
References
-
- Payne S. Family coronaviridae. Viruses. 2017 doi: 10.1016/B978-0-12-803109-4.00017-9. - DOI
-
- Sahin AR, et al. Novel coronavirus (COVID-19) outbreak: a review of the current literature. EJMO. 2020;4:1–7.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

