Biosynthesis and Mechanism of Action of the Cell Wall Targeting Antibiotic Hypeptin
- PMID: 33768646
- PMCID: PMC8252469
- DOI: 10.1002/anie.202102224
Biosynthesis and Mechanism of Action of the Cell Wall Targeting Antibiotic Hypeptin
Abstract
Hypeptin is a cyclodepsipeptide antibiotic produced by Lysobacter sp. K5869, isolated from an environmental sample by the iChip technology, dedicated to the cultivation of previously uncultured microorganisms. Hypeptin shares structural features with teixobactin and exhibits potent activity against a broad spectrum of gram-positive pathogens. Using comprehensive in vivo and in vitro analyses, we show that hypeptin blocks bacterial cell wall biosynthesis by binding to multiple undecaprenyl pyrophosphate-containing biosynthesis intermediates, forming a stoichiometric 2:1 complex. Resistance to hypeptin did not readily develop in vitro. Analysis of the hypeptin biosynthetic gene cluster (BGC) supported a model for the synthesis of the octapeptide. Within the BGC, two hydroxylases were identified and characterized, responsible for the stereoselective β-hydroxylation of four building blocks when bound to peptidyl carrier proteins. In vitro hydroxylation assays corroborate the biosynthetic hypothesis and lead to the proposal of a refined structure for hypeptin.
Keywords: antibiotic; cell wall; cyclodepsipeptide; hydroxylase; lipid II.
© 2021 The Authors. Angewandte Chemie International Edition published by Wiley-VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



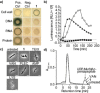

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

