Down-regulation of STAT3 enhanced chemokine expression and neutrophil recruitment in biliary atresia
- PMID: 33769466
- PMCID: PMC8035628
- DOI: 10.1042/CS20201366
Down-regulation of STAT3 enhanced chemokine expression and neutrophil recruitment in biliary atresia
Abstract
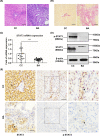
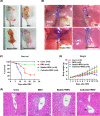
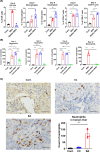
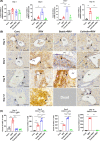
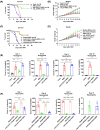
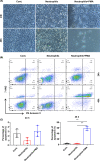
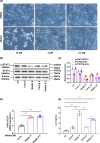
Biliary atresia (BA) is an immune-related disorder and signal transducer and activator of transcription 3 (STAT3) is a key signalling molecule in inflammation. The present study was designed to clarify the function of STAT3 in BA. STAT3 expression was examined in patients and a mouse BA model in which STAT3 levels were further altered with a specific inhibitor or activator. Neutrophil accumulation and the levels of the neutrophil chemoattractants (C-X-C motif) ligand 1 (CXCL1) and IL-8 were determined. The effects of STAT3 inhibition on IL-8 expression were examined in human biliary epithelial cell (BEC) cultures. Functional changes in liver STAT3+ neutrophils in the mouse model were analysed with 10× single cell RNA-seq methods. Results showed STAT3 and p-STAT3 expression was reduced in BA liver tissue compared with control samples. Administration of a STAT3 inhibitor increased jaundice and mortality and reduced body weight in BA mice. In contrast, the STAT3 activator ameliorated BA symptoms. Extensive neutrophil accumulation together with CXCL1 up-regulation, both of which were suppressed by an anti-CXCL1 antibody, were observed in the STAT3 inhibitor-treated group. Recombinant IL-8 administration increased disease severity in BA mice, and the STAT3 activator had the reverse effect. Inhibiting STAT3 increased apoptosis of human BECs together with up-regulated IL-8 expression. RNA-seq analysis revealed reduced the numbers of STAT3 expressing neutrophil in BA which was accompanied by marked enhanced interferon-related antiviral activities. In conclusion, STAT3 reduction, enhanced IL-8 and CXCL1 expression and promoted the accumulation of interferon-responsive neutrophils resulting in BEC damage in BA.
Keywords: Biliary atresia; Neutrophil chemoattractants; STAT3.
© 2021 The Author(s).
Conflict of interest statement
The authors declare that there are no competing interests associated with the manuscript.
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

