Quiescin sulfhydryl oxidase 1 promotes sorafenib-induced ferroptosis in hepatocellular carcinoma by driving EGFR endosomal trafficking and inhibiting NRF2 activation
- PMID: 33770521
- PMCID: PMC8024711
- DOI: 10.1016/j.redox.2021.101942
Quiescin sulfhydryl oxidase 1 promotes sorafenib-induced ferroptosis in hepatocellular carcinoma by driving EGFR endosomal trafficking and inhibiting NRF2 activation
Abstract
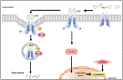
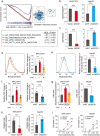
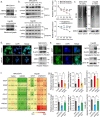
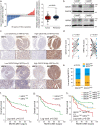
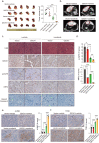
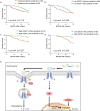
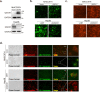
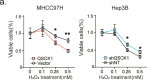
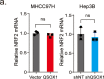
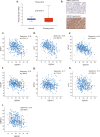
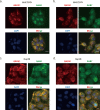
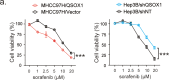
Sorafenib is a first-line molecular-target drug for advanced hepatocellular carcinoma (HCC), but its clinical effects are still limited. In this study we identify Quiescin sulfhydryl oxidase 1 (QSOX1) acting as a cellular pro-oxidant, specifically in the context of sorafenib treatment of HCC. QSOX1 disrupts redox homoeostasis and sensitizes HCC cells to oxidative stress by inhibiting activation of the master antioxidant transcription factor NRF2. A negative correlation between QSOX1 and NRF2 expression was validated in tumor tissues from 151 HCC patients. Mechanistically, QSOX1 restrains EGF-induced EGFR activation by promoting ubiquitination-mediated degradation of EGFR and accelerating its intracellular endosomal trafficking, leading to suppression of NRF2 activity. Additionally, QSOX1 potentiates sorafenib-induced ferroptosis by suppressing NRF2 in vitro and in vivo. In conclusion, the data presented identify QSOX1 as a novel candidate target for sorafenib-based combination therapeutic strategies in HCC or other EGFR-dependent tumor types.
Keywords: Antioxidant; Ferroptosis; HCC; QSOX1; ROS.
Copyright © 2021 The Author(s). Published by Elsevier B.V. All rights reserved.
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
The authors declare no conflict of interest.
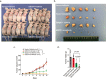
Figures




Similar articles
-
GSTZ1 sensitizes hepatocellular carcinoma cells to sorafenib-induced ferroptosis via inhibition of NRF2/GPX4 axis.Cell Death Dis. 2021 Apr 30;12(5):426. doi: 10.1038/s41419-021-03718-4. Cell Death Dis. 2021. PMID: 33931597 Free PMC article.
-
Tiliroside targets TBK1 to induce ferroptosis and sensitize hepatocellular carcinoma to sorafenib.Phytomedicine. 2023 Mar;111:154668. doi: 10.1016/j.phymed.2023.154668. Epub 2023 Jan 15. Phytomedicine. 2023. PMID: 36657316
-
Camptothecin Sensitizes Hepatocellular Carcinoma Cells to Sorafenib- Induced Ferroptosis Via Suppression of Nrf2.Inflammation. 2023 Aug;46(4):1493-1511. doi: 10.1007/s10753-023-01823-4. Epub 2023 May 12. Inflammation. 2023. PMID: 37171695 Free PMC article.
-
From synergy to resistance: Navigating the complex relationship between sorafenib and ferroptosis in hepatocellular carcinoma.Biomed Pharmacother. 2024 Jan;170:116074. doi: 10.1016/j.biopha.2023.116074. Epub 2023 Dec 25. Biomed Pharmacother. 2024. PMID: 38147732 Review.
-
Roles and Mechanisms of Ferroptosis in Sorafenib Resistance for Hepatocellular Carcinoma.J Hepatocell Carcinoma. 2024 Dec 19;11:2493-2504. doi: 10.2147/JHC.S500084. eCollection 2024. J Hepatocell Carcinoma. 2024. PMID: 39717509 Free PMC article. Review.
Cited by
-
Aberrant m5C hypermethylation mediates intrinsic resistance to gefitinib through NSUN2/YBX1/QSOX1 axis in EGFR-mutant non-small-cell lung cancer.Mol Cancer. 2023 May 9;22(1):81. doi: 10.1186/s12943-023-01780-4. Mol Cancer. 2023. PMID: 37161388 Free PMC article.
-
Identification of PIF1 as a Ferroptosis-Related Prognostic Biomarker Correlated with Immune Infiltration in Hepatocellular Carcinoma.Appl Biochem Biotechnol. 2025 May;197(5):3311-3331. doi: 10.1007/s12010-024-05161-5. Epub 2025 Jan 31. Appl Biochem Biotechnol. 2025. PMID: 39888492
-
AP5Z1 affects hepatocellular carcinoma growth and autophagy by regulating PTEN ubiquitination and modulating the PI3K/Akt/mTOR pathway.J Transl Med. 2025 May 20;23(1):564. doi: 10.1186/s12967-025-06537-9. J Transl Med. 2025. PMID: 40394639 Free PMC article.
-
Mechanisms and therapeutic targets of ErbB family receptors in hepatocellular carcinoma: a narrative review.Transl Cancer Res. 2024 Jun 30;13(6):3156-3178. doi: 10.21037/tcr-24-837. Epub 2024 Jun 27. Transl Cancer Res. 2024. PMID: 38988928 Free PMC article. Review.
-
Ferroptosis in Renal Cancer Therapy: A Narrative Review of Drug Candidates.Cancers (Basel). 2024 Sep 11;16(18):3131. doi: 10.3390/cancers16183131. Cancers (Basel). 2024. PMID: 39335103 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

