Group V Secretory Phospholipase A2 Regulates Endocytosis of Acetylated LDL by Transcriptional Activation of PGK1 in RAW264.7 Macrophage Cell Line
- PMID: 33775979
- PMCID: PMC9135649
- DOI: 10.5551/jat.62216
Group V Secretory Phospholipase A2 Regulates Endocytosis of Acetylated LDL by Transcriptional Activation of PGK1 in RAW264.7 Macrophage Cell Line
Abstract
Aims: It was suggested that group V secretory phospholipase A2 (sPLA2-V) existed in the nucleus. This study examined whether nuclear sPLA2-V plays a role in endocytosis of acetylated low-density lipoprotein (AcLDL) in monocyte/macrophage-like cell line RAW264.7 cells.
Methods: RAW264.7 cells were transfected with shRNA vector targeting sPLA2-V (sPLA2-V-knockdown [KD] cells) or empty vector (sPLA2-V-wild-type [WT] cells). AcLDL endocytosis was assessed by incubation with 125I-AcLDL or AcLDL conjugated with pHrodo. Actin polymerization was assessed by flow cytometry using Alexa Fluor 546-phalloidin.
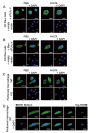
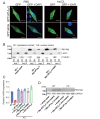
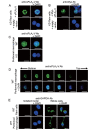
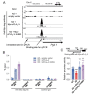
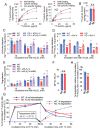
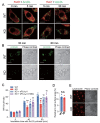
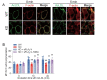
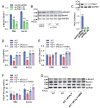
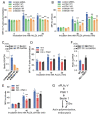
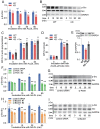
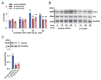
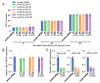
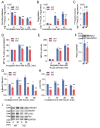
Results: In immunofluorescence microscopic studies, sPLA2-V was detected in the nucleus. ChIP-Seq and ChIP-qPCR analyses showed binding of sPLA2-V to the promoter region of the phosphoglycerate kinase 1 (Pgk1) gene. In the promoter assay, sPLA2-V-KD cells had lower promoter activity of the Pgk1 gene than sPLA2-V-WT cells, and this decrease could be reversed by transfection with a vector encoding sPLA2-V-H48Q that lacks enzymatic activity. Compared with sPLA2-V-WT cells, sPLA2-V-KD cells had decreased PGK1 protein expression, beclin 1 (Beclin1) phosphorylation at S30, and class III PI3-kinase activity that could also be restored by transfection with sPLA2-V-H48Q. sPLA2-V-KD cells had impaired actin polymerization and endocytosis, which was reversed by introduction of sPLA2-V-H48Q or PGK1 overexpression. In sPLA2-V-WT cells, siRNA-mediated depletion of PGK1 suppressed Beclin1 phosphorylation and impaired actin polymerization and intracellular trafficking of pHrodo-conjugated AcLDL.
Conclusions: Nuclear sPLA2-V binds to the Pgk1 gene promoter region and increases its transcriptional activity. sPLA2-V regulates AcLDL endocytosis through PGK1-Beclin1 in a manner that is independent of its enzymatic activity in RAW264.7 cells.
Keywords: Actin polymerization; Endocytosis; Group V secretory phospholipase A2; Macrophage; Phosphoglycerate kinase 1.
Figures
Similar articles
-
Group V secretory phospholipase A2-modified low density lipoprotein promotes foam cell formation by a SR-A- and CD36-independent process that involves cellular proteoglycans.J Biol Chem. 2005 Sep 23;280(38):32746-52. doi: 10.1074/jbc.M502067200. Epub 2005 Jul 21. J Biol Chem. 2005. PMID: 16040605
-
Phosphoglycerate Kinase 1 Phosphorylates Beclin1 to Induce Autophagy.Mol Cell. 2017 Mar 2;65(5):917-931.e6. doi: 10.1016/j.molcel.2017.01.027. Epub 2017 Feb 23. Mol Cell. 2017. PMID: 28238651 Free PMC article.
-
Statin inhibits the expression of secretory phospholipase A2 and subsequent monocyte chemoattractant protein-1 in human endothelial cells.J Cardiovasc Pharmacol. 2014 Dec;64(6):489-96. doi: 10.1097/FJC.0000000000000147. J Cardiovasc Pharmacol. 2014. PMID: 25490414
-
MMP production in human fibrosarcoma cells and their invasiveness are regulated by group IB secretory phospholipase A2 receptor-mediated activation of cytosolic phospholipase A2.Front Biosci. 2008 Jan 1;13:1917-25. doi: 10.2741/2811. Front Biosci. 2008. PMID: 17981679 Review.
-
Group IID, IIE, IIF and III secreted phospholipase A2s.Biochim Biophys Acta Mol Cell Biol Lipids. 2019 Jun;1864(6):803-818. doi: 10.1016/j.bbalip.2018.08.014. Epub 2018 Aug 31. Biochim Biophys Acta Mol Cell Biol Lipids. 2019. PMID: 30905347 Free PMC article. Review.
Cited by
-
Group V Phospholipase A2 Mediates Endothelial Dysfunction and Acute Lung Injury Caused by Methicillin-Resistant Staphylococcus Aureus.Cells. 2021 Jul 8;10(7):1731. doi: 10.3390/cells10071731. Cells. 2021. PMID: 34359901 Free PMC article.
References
-
- Bingham CO 3rd, Fijneman RJ, Friend DS, Goddeau RP, Rogers RA, Austen KF, Arm JP: Low molecular weight group IIA and group V phospholipase A(2) enzymes have different intracellular locations in mouse bone marrow-derived mast cells. J Biol Chem, 1999; 274: 31476-31484 - PubMed
-
- Nardicchi V, Macchioni L, Ferrini M, Goracci G: The presence of a secretory phospholipase A2 in the nuclei of neuronal and glial cells of rat brain cortex. Biochim Biophys Acta, 2007; 1771: 1345-1352 - PubMed
-
- Fayard JM, Tessier C, Pageaux JF, Lagarde M, Laugier C: Nuclear location of PLA2-I in proliferative cells. J Cell Sci, 1998; 111 (Pt 7): 985-994 - PubMed
-
- Oishi T, Tamiya-Koizumi K, Kudo I, Iino S, Takagi K, Yoshida S: Purification and characterization of nuclear alkaline phospholipase A2 in rat ascites hepatoma cells. FEBS Lett, 1996; 394: 55-60 - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

