Reconstruction of ARNT PAS-B Unfolding Pathways by Steered Molecular Dynamics and Artificial Neural Networks
- PMID: 33780250
- PMCID: PMC8047803
- DOI: 10.1021/acs.jctc.0c01308
Reconstruction of ARNT PAS-B Unfolding Pathways by Steered Molecular Dynamics and Artificial Neural Networks
Abstract
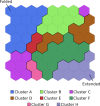
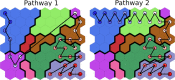
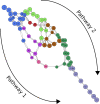
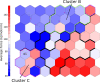
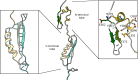
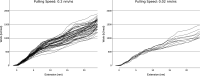
Several experimental studies indicated that large conformational changes, including partial domain unfolding, have a role in the functional mechanisms of the basic helix loop helix Per/ARNT/SIM (bHLH-PAS) transcription factors. Recently, single-molecule atomic force microscopy (AFM) revealed two distinct pathways for the mechanical unfolding of the ARNT PAS-B. In this work we used steered molecular dynamics simulations to gain new insights into this process at an atomistic level. To reconstruct and classify pathways sampled in multiple simulations, we designed an original approach based on the use of self-organizing maps (SOMs). This led us to identify two types of unfolding pathways for the ARNT PAS-B, which are in good agreement with the AFM findings. Analysis of average forces mapped on the SOM revealed a stable conformation of the PAS-B along one pathway, which represents a possible structural model for the intermediate state detected by AFM. The approach here proposed will facilitate the study of other signal transmission mechanisms involving the folding/unfolding of PAS domains.
Conflict of interest statement
The authors declare no competing financial interest.
Figures
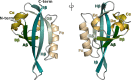


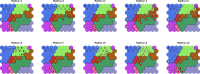
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

