Using the antibody-antigen binding interface to train image-based deep neural networks for antibody-epitope classification
- PMID: 33780441
- PMCID: PMC8032195
- DOI: 10.1371/journal.pcbi.1008864
Using the antibody-antigen binding interface to train image-based deep neural networks for antibody-epitope classification
Abstract
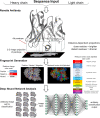
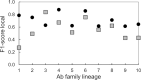
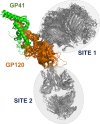
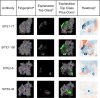
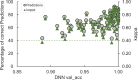
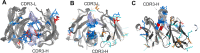
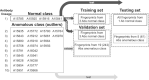
High-throughput B-cell sequencing has opened up new avenues for investigating complex mechanisms underlying our adaptive immune response. These technological advances drive data generation and the need to mine and analyze the information contained in these large datasets, in particular the identification of therapeutic antibodies (Abs) or those associated with disease exposure and protection. Here, we describe our efforts to use artificial intelligence (AI)-based image-analyses for prospective classification of Abs based solely on sequence information. We hypothesized that Abs recognizing the same part of an antigen share a limited set of features at the binding interface, and that the binding site regions of these Abs share share common structure and physicochemical property patterns that can serve as a "fingerprint" to recognize uncharacterized Abs. We combined large-scale sequence-based protein-structure predictions to generate ensembles of 3-D Ab models, reduced the Ab binding interface to a 2-D image (fingerprint), used pre-trained convolutional neural networks to extract features, and trained deep neural networks (DNNs) to classify Abs. We evaluated this approach using Ab sequences derived from human HIV and Ebola viral infections to differentiate between two Abs, Abs belonging to specific B-cell family lineages, and Abs with different epitope preferences. In addition, we explored a different type of DNN method to detect one class of Abs from a larger pool of Abs. Testing on Ab sets that had been kept aside during model training, we achieved average prediction accuracies ranging from 71-96% depending on the complexity of the classification task. The high level of accuracies reached during these classification tests suggests that the DNN models were able to learn a series of structural patterns shared by Abs belonging to the same class. The developed methodology provides a means to apply AI-based image recognition techniques to analyze high-throughput B-cell sequencing datasets (repertoires) for Ab classification.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




References
-
- Dati F, Schumann G, Thomas L, Aguzzi F, Baudner S, Bienvenu J, et al. Consensus of a group of professional societies and diagnostic companies on guidelines for interim reference ranges for 14 proteins in serum based on the standardization against the IFCC/BCR/CAP Reference Material (CRM 470). International Federation of Clinical Chemistry, Community Bureau of Reference of the Commission of the European Communities, College of American Pathologists. Eur J Clin Chem Clin Biochem 1996; 34(6):517–20. ed2020. - PubMed
-
- Briggs AW, Goldfless SJ, Timberlake S, Belmont BJ, Clouser CR, Koppstein D, et al. Tumor-infiltrating immune repertoires captured by single-cell barcoding in emulsion. bioRxiv [Preprint]. 2017. bioRxiv 13841. Available from: 10.1101/134841. - DOI
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

