A protein structural study based on the centrality analysis of protein sequence feature networks
- PMID: 33780482
- PMCID: PMC8006989
- DOI: 10.1371/journal.pone.0248861
A protein structural study based on the centrality analysis of protein sequence feature networks
Abstract
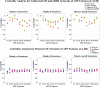
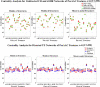
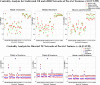
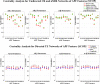
In this paper, we use network approaches to analyze the relations between protein sequence features for the top hierarchical classes of CATH and SCOP. We use fundamental connectivity measures such as correlation (CR), normalized mutual information rate (nMIR), and transfer entropy (TE) to analyze the pairwise-relationships between the protein sequence features, and use centrality measures to analyze weighted networks constructed from the relationship matrices. In the centrality analysis, we find both commonalities and differences between the different protein 3D structural classes. Results show that all top hierarchical classes of CATH and SCOP present strong non-deterministic interactions for the composition and arrangement features of Cystine (C), Methionine (M), Tryptophan (W), and also for the arrangement features of Histidine (H). The different protein 3D structural classes present different preferences in terms of their centrality distributions and significant features.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures








References
-
- Wang J, Wang Z, Tian X. Bioinformatics: Fundementals and applications. Tsinghua University Press. 2014.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

