Comparative genomics of Chlamydomonas
- PMID: 33793842
- PMCID: PMC8226300
- DOI: 10.1093/plcell/koab026
Comparative genomics of Chlamydomonas
Abstract
Despite its role as a reference organism in the plant sciences, the green alga Chlamydomonas reinhardtii entirely lacks genomic resources from closely related species. We present highly contiguous and well-annotated genome assemblies for three unicellular C. reinhardtii relatives: Chlamydomonas incerta, Chlamydomonas schloesseri, and the more distantly related Edaphochlamys debaryana. The three Chlamydomonas genomes are highly syntenous with similar gene contents, although the 129.2 Mb C. incerta and 130.2 Mb C. schloesseri assemblies are more repeat-rich than the 111.1 Mb C. reinhardtii genome. We identify the major centromeric repeat in C. reinhardtii as a LINE transposable element homologous to Zepp (the centromeric repeat in Coccomyxa subellipsoidea) and infer that centromere locations and structure are likely conserved in C. incerta and C. schloesseri. We report extensive rearrangements, but limited gene turnover, between the minus mating type loci of these Chlamydomonas species. We produce an eight-species core-Reinhardtinia whole-genome alignment, which we use to identify several hundred false positive and missing genes in the C. reinhardtii annotation and >260,000 evolutionarily conserved elements in the C. reinhardtii genome. In summary, these resources will enable comparative genomics analyses for C. reinhardtii, significantly extending the analytical toolkit for this emerging model system.
� The Author(s) 2021. Published by Oxford University Press on behalf of American Society of Plant Biologists.
Figures





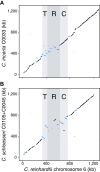




References
-
- Blaby IK, Blaby-Haas CE (2017) Genomics and functional genomics in Chlamydomonas reinhardtii. In Hippler M, ed., Chlamydomonas: Molecular Genetics and Physiology. Springer, New York
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

