Telomere Length Dynamics and Chromosomal Instability for Predicting Individual Radiosensitivity and Risk via Machine Learning
- PMID: 33800260
- PMCID: PMC8002073
- DOI: 10.3390/jpm11030188
Telomere Length Dynamics and Chromosomal Instability for Predicting Individual Radiosensitivity and Risk via Machine Learning
Abstract
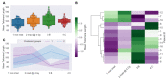
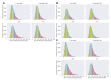
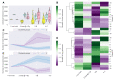
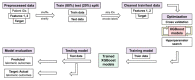
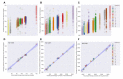
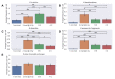
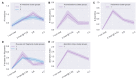
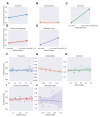
The ability to predict a cancer patient's response to radiotherapy and risk of developing adverse late health effects would greatly improve personalized treatment regimens and individual outcomes. Telomeres represent a compelling biomarker of individual radiosensitivity and risk, as exposure can result in dysfunctional telomere pathologies that coincidentally overlap with many radiation-induced late effects, ranging from degenerative conditions like fibrosis and cardiovascular disease to proliferative pathologies like cancer. Here, telomere length was longitudinally assessed in a cohort of fifteen prostate cancer patients undergoing Intensity Modulated Radiation Therapy (IMRT) utilizing Telomere Fluorescence in situ Hybridization (Telo-FISH). To evaluate genome instability and enhance predictions for individual patient risk of secondary malignancy, chromosome aberrations were assessed utilizing directional Genomic Hybridization (dGH) for high-resolution inversion detection. We present the first implementation of individual telomere length data in a machine learning model, XGBoost, trained on pre-radiotherapy (baseline) and in vitro exposed (4 Gy γ-rays) telomere length measurements, to predict post radiotherapy telomeric outcomes, which together with chromosomal instability provide insight into individual radiosensitivity and risk for radiation-induced late effects.
Keywords: IMRT; chromosomal instability; individual radiosensitivity; inversions; late effects; machine learning; personalized medicine; prostate cancer; telomeres.
Conflict of interest statement
S.M.B. is a cofounder and scientific advisory board member of KromaTiD, Inc.
Figures


Similar articles
-
The role of telomeres in predicting individual radiosensitivity of patients with cancer in the era of personalized radiotherapy.Cancer Treat Rev. 2015 Apr;41(4):354-60. doi: 10.1016/j.ctrv.2015.02.005. Epub 2015 Feb 11. Cancer Treat Rev. 2015. PMID: 25704912 Review.
-
Long but dysfunctional telomeres correlate with chromosomal radiosensitivity in a mouse AML cell line.Int J Radiat Biol. 2001 Dec;77(12):1151-62. doi: 10.1080/09553000110075220. Int J Radiat Biol. 2001. PMID: 11747540
-
Comparison of Individual Radiosensitivity to γ-Rays and Carbon Ions.Front Oncol. 2016 Jun 13;6:137. doi: 10.3389/fonc.2016.00137. eCollection 2016. Front Oncol. 2016. PMID: 27379201 Free PMC article.
-
Telomeres and NextGen CO-FISH: Directional Genomic Hybridization (Telo-dGH™).Methods Mol Biol. 2017;1587:103-112. doi: 10.1007/978-1-4939-6892-3_10. Methods Mol Biol. 2017. PMID: 28324502
-
Using telomeric chromosomal aberrations to evaluate clastogen-induced genomic instability in mammalian cells.Chromosome Res. 2020 Dec;28(3-4):259-276. doi: 10.1007/s10577-020-09641-2. Epub 2020 Sep 17. Chromosome Res. 2020. PMID: 32940874 Review.
Cited by
-
A correlation graph attention network for classifying chromosomal instabilities from histopathology whole-slide images.iScience. 2023 May 18;26(6):106874. doi: 10.1016/j.isci.2023.106874. eCollection 2023 Jun 16. iScience. 2023. PMID: 37260749 Free PMC article.
-
Machine Learning & Molecular Radiation Tumor Biomarkers.Semin Radiat Oncol. 2023 Jul;33(3):243-251. doi: 10.1016/j.semradonc.2023.03.002. Semin Radiat Oncol. 2023. PMID: 37331779 Free PMC article. Review.
-
Telomeric RNA (TERRA) increases in response to spaceflight and high-altitude climbing.Commun Biol. 2024 Jun 11;7(1):698. doi: 10.1038/s42003-024-06014-x. Commun Biol. 2024. PMID: 38862827 Free PMC article.
-
Role of telomere length in human carcinogenesis (Review).Int J Oncol. 2023 Jul;63(1):78. doi: 10.3892/ijo.2023.5526. Epub 2023 May 26. Int J Oncol. 2023. PMID: 37232367 Free PMC article. Review.
-
Radiation Biomarkers: Silver Bullet, or Wild Goose Chase?J Pers Med. 2021 Jun 25;11(7):603. doi: 10.3390/jpm11070603. J Pers Med. 2021. PMID: 34202274 Free PMC article.
References
-
- Carver J.R., Shapiro C.L., Ng A., Jacobs L., Schwartz C., Virgo K.S., Hagerty K.L., Somerfield M.R., Vaughn D.J., ASCO Cancer Survivorship Expert Panel American Society of Clinical Oncology Clinical Evidence Review on the Ongoing Care of Adult Cancer Survivors: Cardiac and Pulmonary Late Effects. J. Clin. Oncol. 2007;25:3991–4008. doi: 10.1200/JCO.2007.10.9777. - DOI - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

