Isolation and Characterization of a Novel Lytic Bacteriophage against the K2 Capsule-Expressing Hypervirulent Klebsiella pneumoniae Strain 52145, and Identification of Its Functional Depolymerase
- PMID: 33801047
- PMCID: PMC8003838
- DOI: 10.3390/microorganisms9030650
Isolation and Characterization of a Novel Lytic Bacteriophage against the K2 Capsule-Expressing Hypervirulent Klebsiella pneumoniae Strain 52145, and Identification of Its Functional Depolymerase
Abstract
Klebsiella pneumoniae is among the leading bacteria that cause nosocomial infections. The capsule of this Gram-negative bacterium is a dominant virulence factor, with a prominent role in defense and biofilm formation. Bacteriophages, which are specific for one bacterial strain and its capsule type, can evoke the lysis of bacterial cells, aided by polysaccharide depolymerase enzymes. In this study, we isolated and characterized a bacteriophage against the nosocomial K. pneumoniae 52145 strain with K2 capsular serotype. The phage showed a narrow host range and stable lytic activity, even when exposed to different temperatures or detergents. Preventive effect of the phage in a nasal colonization model was investigated in vivo. Phlyogenetic analysis showed that the newly isolated Klebsiella phage B1 belongs to the Webervirus genus in Drexlerviridae family. We identified the location of the capsule depolymerase gene of the new phage, which was amplified, cloned, expressed, and purified. The efficacy of the recombinant B1dep depolymerase was tested by spotting on K. pneumoniae strains and it was confirmed that the extract lowers the thickness of the bacterium lawn as it degrades the protective capsule on bacterial cells. As K. pneumoniae strains possessing the K2 serotype have epidemiological importance, the B1 phage and its depolymerase are promising candidates for use as possible antimicrobial agents.
Keywords: K2 serotype; Klebsiella pneumoniae; bacteriophage; capsule depolymerase; tail fibers.
Conflict of interest statement
The authors declare that there is no conflict of interest regarding the publication of this paper.
Figures
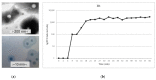
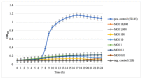


 ).
).

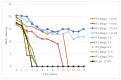
References
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

