Distribution of Extended-Spectrum β-Lactamase (ESBL)-Encoding Genes among Multidrug-Resistant Gram-Negative Pathogens Collected from Three Different Countries
- PMID: 33801418
- PMCID: PMC7998439
- DOI: 10.3390/antibiotics10030247
Distribution of Extended-Spectrum β-Lactamase (ESBL)-Encoding Genes among Multidrug-Resistant Gram-Negative Pathogens Collected from Three Different Countries
Abstract
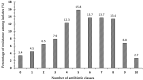
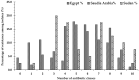
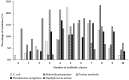
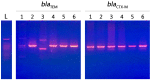
The incidence of Extended-spectrum β-lactamase (ESBL)-encoding genes (blaCTX-M and blaTEM) among Gram-negative multidrug-resistant pathogens collected from three different countries was investigated. Two hundred and ninety-two clinical isolates were collected from Egypt (n = 90), Saudi Arabia (n = 162), and Sudan (n = 40). Based on the antimicrobial sensitivity against 20 antimicrobial agents from 11 antibiotic classes, the most resistant strains were selected and identified using the Vitek2 system and 16S rRNA gene sequence analysis. A total of 85.6% of the isolates were found to be resistant to more than three antibiotic classes. The ratios of the multidrug-resistant strains for Egypt, Saudi Arabia, and Sudan were 74.4%, 90.1%, and 97.5%, respectively. Escherichia coli, Klebsiella pneumoniae, and Pseudomonas aeruginosa showed inconstant resistance levels to the different classes of antibiotics. Escherichia coli and Klebsiella pneumoniae had the highest levels of resistance against macrolides followed by penicillins and cephalosporin, while Pseudomonas aeruginosa was most resistant to penicillins followed by classes that varied among different countries. The isolates were positive for the presence of the blaCTX-M and blaTEM genes. The blaCTX-M gene was the predominant gene in all isolates (100%), while blaTEM was detected in 66.7% of the selected isolates. This work highlights the detection of multidrug-resistant bacteria and resistant genes among different countries. We suggest that the medical authorities urgently implement antimicrobial surveillance plans and infection control policies for early detection and effective prevention of the rapid spread of these pathogens.
Keywords: antimicrobial resistance; clinical samples; different countries; multidrug-resistant pathogens; resistant-genes.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Antimicrobial resistance profiles of Pseudomonas aeruginosa, Escherichia coli and Klebsiella pneumoniae strains isolated from broiler chickens.Food Microbiol. 2024 Jun;120:104476. doi: 10.1016/j.fm.2024.104476. Epub 2024 Jan 10. Food Microbiol. 2024. PMID: 38431322
-
Extended-spectrum beta-lactamase-producing Escherichia coli and Klebsiella pneumoniae: insights from a tertiary hospital in Southern Thailand.Microbiol Spectr. 2024 Jul 2;12(7):e0021324. doi: 10.1128/spectrum.00213-24. Epub 2024 May 29. Microbiol Spectr. 2024. PMID: 38809095 Free PMC article.
-
Extended Spectrum Beta Lactamase (ESBL), blaTEM,blaSHV and blaCTX-M, Resistance Genes in Community and Healthcare Associated Gram Negative Bacteria from Osun State, Nigeria.Infect Disord Drug Targets. 2021;21(4):595-602. doi: 10.2174/1871526520999200729181559. Infect Disord Drug Targets. 2021. PMID: 32729432
-
Phenotypic and Genotypic Characterization of Extended Spectrum Beta-Lactamase-Producing Clinical Isolates of Escherichia coli and Klebsiella pneumoniae in Two Kenyan Facilities: A National Referral and a Level Five Hospital.Int J Microbiol. 2024 Feb 14;2024:7463899. doi: 10.1155/2024/7463899. eCollection 2024. Int J Microbiol. 2024. PMID: 38384586 Free PMC article.
-
Retrospective Analysis on Antimicrobial Resistance Trends and Prevalence of β-lactamases in Escherichia coli and ESKAPE Pathogens Isolated from Arabian Patients during 2000-2020.Microorganisms. 2020 Oct 21;8(10):1626. doi: 10.3390/microorganisms8101626. Microorganisms. 2020. PMID: 33096921 Free PMC article. Review.
Cited by
-
Antimicrobial Resistance Patterns and Risk Factors Associated with ESBL-Producing and MDR Escherichia coli in Hospital and Environmental Settings in Lusaka, Zambia: Implications for One Health, Antimicrobial Stewardship and Surveillance Systems.Microorganisms. 2023 Jul 31;11(8):1951. doi: 10.3390/microorganisms11081951. Microorganisms. 2023. PMID: 37630511 Free PMC article.
-
Whole Genome Characterization of the High-Risk Clone ST383 Klebsiella pneumoniae with a Simultaneous Carriage of blaCTX-M-14 on IncL/M Plasmid and blaCTX-M-15 on Convergent IncHI1B/IncFIB Plasmid from Egypt.Microorganisms. 2022 May 26;10(6):1097. doi: 10.3390/microorganisms10061097. Microorganisms. 2022. PMID: 35744615 Free PMC article.
-
A Multi-Point Surveillance for Antimicrobial Resistance Profiles among Clinical Isolates of Gram-Negative Bacteria Recovered from Major Ha'il Hospitals, Saudi Arabia.Microorganisms. 2021 Sep 24;9(10):2024. doi: 10.3390/microorganisms9102024. Microorganisms. 2021. PMID: 34683344 Free PMC article.
-
Phenotypic and molecular characterization of extended spectrum- and metallo- beta lactamase producing Pseudomonas aeruginosa clinical isolates from Egypt.Infection. 2024 Dec;52(6):2399-2414. doi: 10.1007/s15010-024-02297-8. Epub 2024 Jun 2. Infection. 2024. PMID: 38824475 Free PMC article.
-
Genetic Analysis, Population Structure, and Characterisation of Multidrug-Resistant Klebsiella pneumoniae from the Al-Hofuf Region of Saudi Arabia.Pathogens. 2021 Aug 28;10(9):1097. doi: 10.3390/pathogens10091097. Pathogens. 2021. PMID: 34578130 Free PMC article.
References
-
- Sahm D.F., Thornsberry C., Mayfield D.C., Jones M.E., Karlowsky J.A. Multidrug-resistant urinary tract isolates of Escherichia coli: Prevalence and patient demographics in the United States in 2000. Antimicrob. Agents Chemother. 2001;45:1402–1406. doi: 10.1128/AAC.45.5.1402-1406.2001. - DOI - PMC - PubMed
-
- Karlowsky J.A., Jones M.E., Draghi D.C., Thornsberry C., Sahm D.F., Volturo G.A. Prevalence and antimicrobial susceptibilities of bacteria isolated from blood cultures of hospitalized patients in the United States in 2002. Ann. Clin. Microbiol. Antimicrob. 2004;3:7. doi: 10.1186/1476-0711-3-7. - DOI - PMC - PubMed
-
- Van De Sande-Bruinsma N., Grundmann H., Verloo D., Tiemersma E., Monen J., Goossens H., Ferech M., European Antimicrobial Resistance Surveillance System Group. European Surveillance of Antimicrobial Consumption Project Group Antimicrobial drug use and resistance in Europe. Emerg. Infect. Dis. 2008;14:1722–1730. doi: 10.3201/eid1411.070467. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

