Different Stages of Quiescence, Senescence, and Cell Stress Identified by Molecular Algorithm Based on the Expression of Ki67, RPS6, and Beta-Galactosidase Activity
- PMID: 33803589
- PMCID: PMC8002939
- DOI: 10.3390/ijms22063102
Different Stages of Quiescence, Senescence, and Cell Stress Identified by Molecular Algorithm Based on the Expression of Ki67, RPS6, and Beta-Galactosidase Activity
Abstract
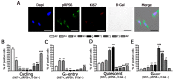
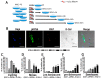
During their life span, cells have two possible states: a non-cycling, quiescent state (G0) and a cycling, activated state. Cells may enter a reversible G0 state of quiescence or, alternatively, they may undergo an irreversible G0 state. The latter may be a physiological differentiation or, following a stress event, a senescent status. Discrimination among the several G0 states represents a significant investigation, since quiescence, differentiation, and senescence are progressive phenomena with intermediate transitional stages. We used the expression of Ki67, RPS6, and beta-galactosidase to identify healthy cells that progressively enter and leave quiescence through G0-entry, G0 and G0-alert states. We then evaluated how cells may enter senescence following a genotoxic stressful event. We identified an initial stress stage with the expression of beta-galactosidase and Ki67 proliferation marker. Cells may recover from stress events or become senescent passing through early and late senescence states. Discrimination between quiescence and senescence was based on the expression of RPS6, a marker of active protein synthesis that is present in senescent cells but absent in quiescent cells. Even taking into account that fixed G0 states do not exist, our molecular algorithm may represent a method for identifying turning points of G0 transitional states that continuously change.
Keywords: cell cycle; mesenchymal stem cells; quiescence; senescence.
Conflict of interest statement
No conflict of interest is declared.
Figures


References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

