Quantitative Proteomic Approach Reveals Altered Metabolic Pathways in Response to the Inhibition of Lysine Deacetylases in A549 Cells under Normoxia and Hypoxia
- PMID: 33806075
- PMCID: PMC8036653
- DOI: 10.3390/ijms22073378
Quantitative Proteomic Approach Reveals Altered Metabolic Pathways in Response to the Inhibition of Lysine Deacetylases in A549 Cells under Normoxia and Hypoxia
Abstract
Growing evidence is showing that acetylation plays an essential role in cancer, but studies on the impact of KDAC inhibition (KDACi) on the metabolic profile are still in their infancy. Here, we analyzed, by using an iTRAQ-based quantitative proteomics approach, the changes in the proteome of KRAS-mutated non-small cell lung cancer (NSCLC) A549 cells in response to trichostatin-A (TSA) and nicotinamide (NAM) under normoxia and hypoxia. Part of this response was further validated by molecular and biochemical analyses and correlated with the proliferation rates, apoptotic cell death, and activation of ROS scavenging mechanisms in opposition to the ROS production. Despite the differences among the KDAC inhibitors, up-regulation of glycolysis, TCA cycle, oxidative phosphorylation and fatty acid synthesis emerged as a common metabolic response underlying KDACi. We also observed that some of the KDACi effects at metabolic levels are enhanced under hypoxia. Furthermore, we used a drug repositioning machine learning approach to list candidate metabolic therapeutic agents for KRAS mutated NSCLC. Together, these results allow us to better understand the metabolic regulations underlying KDACi in NSCLC, taking into account the microenvironment of tumors related to hypoxia, and bring new insights for the future rational design of new therapies.
Keywords: NSCLC; cancer metabolism; hypoxia; lysine deacetylase inhibitors.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

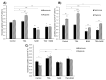

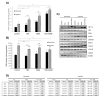

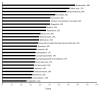
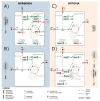

Similar articles
-
A large scale proteome analysis of the gefitinib primary resistance overcome by KDAC inhibition in KRAS mutated adenocarcinoma cells overexpressing amphiregulin.J Proteomics. 2019 Mar 20;195:114-124. doi: 10.1016/j.jprot.2019.01.009. Epub 2019 Jan 17. J Proteomics. 2019. PMID: 30660770
-
Quantitative proteomic approach to understand metabolic adaptation in non-small cell lung cancer.J Proteome Res. 2014 Nov 7;13(11):4695-704. doi: 10.1021/pr500327v. Epub 2014 Aug 25. J Proteome Res. 2014. PMID: 25029028
-
Sensitization to gamma-irradiation-induced cell cycle arrest and apoptosis by the histone deacetylase inhibitor trichostatin A in non-small cell lung cancer (NSCLC) cells.Cancer Biol Ther. 2009 May;8(9):823-31. doi: 10.4161/cbt.8.9.8143. Epub 2009 May 10. Cancer Biol Ther. 2009. PMID: 19270532
-
Targeting metabolic reprogramming in KRAS-driven cancers.Int J Clin Oncol. 2017 Aug;22(4):651-659. doi: 10.1007/s10147-017-1156-4. Epub 2017 Jun 24. Int J Clin Oncol. 2017. PMID: 28647837 Review.
-
Targeting the Cancer Epigenome with Histone Deacetylase Inhibitors in Osteosarcoma.Adv Exp Med Biol. 2020;1258:55-75. doi: 10.1007/978-3-030-43085-6_4. Adv Exp Med Biol. 2020. PMID: 32767234 Review.
Cited by
-
Integrative Analysis of Proteomics and Metabolism Reveals the Potential Roles of Arachidonic Acid Metabolism in Hypoxia Response in Mouse Spleen.Molecules. 2022 Nov 21;27(22):8102. doi: 10.3390/molecules27228102. Molecules. 2022. PMID: 36432203 Free PMC article.
-
Mechanisms of Adipose Tissue Metabolism in Naturally Grazing Sheep at Different Growth Stages: Insights from mRNA and miRNA Profiles.Int J Mol Sci. 2025 Apr 2;26(7):3324. doi: 10.3390/ijms26073324. Int J Mol Sci. 2025. PMID: 40244192 Free PMC article.
-
Hypoxia Changes Energy Metabolism and Growth Rate in Non-Small Cell Lung Cancer Cells.Cancers (Basel). 2023 Apr 26;15(9):2472. doi: 10.3390/cancers15092472. Cancers (Basel). 2023. PMID: 37173939 Free PMC article.
-
Energy metabolism as the hub of advanced non-small cell lung cancer management: a comprehensive view in the framework of predictive, preventive, and personalized medicine.EPMA J. 2024 Apr 8;15(2):289-319. doi: 10.1007/s13167-024-00357-5. eCollection 2024 Jun. EPMA J. 2024. PMID: 38841622 Free PMC article. Review.
-
Evaluation of AMG510 Therapy on KRAS-Mutant Non-Small Cell Lung Cancer and Colorectal Cancer Cell Using a 3D Invasive Tumor Spheroid System under Normoxia and Hypoxia.Bioengineering (Basel). 2022 Dec 12;9(12):792. doi: 10.3390/bioengineering9120792. Bioengineering (Basel). 2022. PMID: 36550998 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

