Absence of 4-Formylaminooxyvinylglycine Production by Pseudomonas fluorescens WH6 Results in Resource Reallocation from Secondary Metabolite Production to Rhizocompetence
- PMID: 33807194
- PMCID: PMC8067088
- DOI: 10.3390/microorganisms9040717
Absence of 4-Formylaminooxyvinylglycine Production by Pseudomonas fluorescens WH6 Results in Resource Reallocation from Secondary Metabolite Production to Rhizocompetence
Abstract
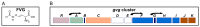
Pseudomonas fluorescens WH6 produces the non-proteinogenic amino acid 4-formylaminooxyvinylglycine (FVG), a secondary metabolite with antibacterial and pre-emergent herbicidal activities. The gvg operon necessary for FVG production encodes eight required genes: one regulatory (gvgR), two of unknown functional potential (gvgA and C), three with putative biosynthetic function (gvgF, H, and I), and two small ORFs (gvgB and G). To gain insight into the role of GvgA and C in FVG production, we compared the transcriptome of knockout (KO) mutants of gvgR, A, and C to wild type (WT) to test two hypotheses: (1) GvgA and GvgC play a regulatory role in FVG production and (2) non-gvg cluster genes are regulated by GvgA and GvgC. Our analyses show that, collectively, 687 genes, including the gvg operon, are differentially expressed in all KO strains versus WT, representing >10% of the genome. Fifty-one percent of these genes were similarly regulated in all KO strains with GvgC having the greatest number of uniquely regulated genes. Additional transcriptome data suggest cluster regulation through feedback of a cluster product. We also discovered that FVG biosynthesis is regulated by L-glu, L-asp, L-gln, and L-asn and that resources are reallocated in KO strains to increase phenotypes involved in rhizocompetence including motility, biofilm formation, and denitrification. Altogether, differential transcriptome analyses of mutants suggest that regulation of the cluster is multifaceted and the absence of FVG production or its downregulation can dramatically shift the lifestyle of WH6.
Keywords: Pseudomonas fluorescens; natural herbicide; regulation of secondary metabolites; vinylglycine.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Figures





References
-
- Keswani C., Singh H.B., García-Estrada C., Caradus J., He Y.W., Mezaache-Aichour S., Glare T.R., Borriss R., Sansinenea E. Antimicrobial secondary metabolites from agriculturally important bacteria as next-generation pesticides. Appl. Microbiol. Biotechnol. 2020;104:1013–1034. doi: 10.1007/s00253-019-10300-8. - DOI - PubMed
-
- Elliott L.F., Azevedo M.D., Mueller-Warrant G.W., Horwath W.R. Weed control with rhizobacteria. Soil Sci. Agrochem. Ecol. 1998;33:3–7.
-
- Halgren A., Azevedo M., Mills D., Armstrong D., Thimmaiah M., Mcphail K., Banowetz G.M. Selective inhibition of Erwinia amylovora by the herbicidally active germination-arrest factor (GAF) produced by Pseudomonas bacteria. J. Appl. Microbiol. 2011;111:949–959. doi: 10.1111/j.1365-2672.2011.05098.x. - DOI - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

