The Arabidopsis thaliana chloroplast division protein FtsZ1 counterbalances FtsZ2 filament stability in vitro
- PMID: 33812992
- PMCID: PMC8142252
- DOI: 10.1016/j.jbc.2021.100627
The Arabidopsis thaliana chloroplast division protein FtsZ1 counterbalances FtsZ2 filament stability in vitro
Abstract
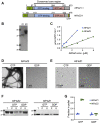
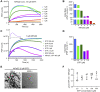
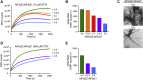
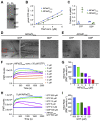
Bacterial cell and chloroplast division are driven by a contractile "Z ring" composed of the tubulin-like cytoskeletal GTPase FtsZ. Unlike bacterial Z rings, which consist of a single FtsZ, the chloroplast Z ring in plants is composed of two FtsZ proteins, FtsZ1 and FtsZ2. Both are required for chloroplast division in vivo, but their biochemical relationship is poorly understood. We used GTPase assays, light scattering, transmission electron microscopy, and sedimentation assays to investigate the assembly behavior of purified Arabidopsis thaliana (At) FtsZ1 and AtFtsZ2 both individually and together. Both proteins exhibited GTPase activity. AtFtsZ2 assembled relatively quickly, forming protofilament bundles that were exceptionally stable, as indicated by their sustained assembly and slow disassembly. AtFtsZ1 did not form detectable protofilaments on its own. When mixed with AtFtsZ2, AtFtsZ1 reduced the extent and rate of AtFtsZ2 assembly, consistent with its previously demonstrated ability to promote protofilament subunit turnover in living cells. Mixing the two FtsZ proteins did not increase the overall GTPase activity, indicating that the effect of AtFtsZ1 on AtFtsZ2 assembly was not due to a stimulation of GTPase activity. However, the GTPase activity of AtFtsZ1 was required to reduce AtFtsZ2 assembly. Truncated forms of AtFtsZ1 and AtFtsZ2 consisting of only their conserved core regions largely recapitulated the behaviors of the full-length proteins. Our in vitro findings provide evidence that FtsZ1 counterbalances the stability of FtsZ2 filaments in the regulation of chloroplast Z-ring dynamics and suggest that restraining FtsZ2 self-assembly is a critical function of FtsZ1 in chloroplasts.
Keywords: Arabidopsis thaliana; FtsZ dynamics; GTPase; chloroplast; cytoskeleton; plant biochemistry.
Copyright © 2021 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Conflict of interest The authors declare that they have no conflicts of interest with the contents of this article.
Figures


References
-
- Gould S.B., Waller R.F., McFadden G.I. Plastid evolution. Annu. Rev. Plant Biol. 2008;59:491–517. - PubMed
-
- Miyagishima S.-Y., Nakanishi H., Kabeya Y. Vol. 291. 2011. pp. 115–153. (Structure, regulation, and evolution of the plastid division machinery. Int. Rev. Cell Mol. Biol.). - PubMed
-
- Osteryoung K.W., Pyke K.A. Division and dynamic morphology of plastids. Annu. Rev. Plant Biol. 2014;65:443–472. - PubMed
-
- Bi E.F., Lutkenhaus J. FtsZ ring structure associated with division in Escherichia coli. Nature. 1991;354:161–164. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

