Molluscan mitochondrial genomes break the rules
- PMID: 33813887
- PMCID: PMC8059616
- DOI: 10.1098/rstb.2020.0159
Molluscan mitochondrial genomes break the rules
Abstract
The first animal mitochondrial genomes to be sequenced were of several vertebrates and model organisms, and the consistency of genomic features found has led to a 'textbook description'. However, a more broad phylogenetic sampling of complete animal mitochondrial genomes has found many cases where these features do not exist, and the phylum Mollusca is especially replete with these exceptions. The characterization of full mollusc mitogenomes required considerable effort involving challenging molecular biology, but has created an enormous catalogue of surprising deviations from that textbook description, including wide variation in size, radical genome rearrangements, gene duplications and losses, the introduction of novel genes, and a complex system of inheritance dubbed 'doubly uniparental inheritance'. Here, we review the extraordinary variation in architecture, molecular functioning and intergenerational transmission of molluscan mitochondrial genomes. Such features represent a great potential for the discovery of biological history, processes and functions that are novel for animal mitochondrial genomes. This provides a model system for studying the evolution and the manifold roles that mitochondria play in organismal physiology, and many ways that the study of mitochondrial genomes are useful for phylogeny and population biology. This article is part of the Theo Murphy meeting issue 'Molluscan genomics: broad insights and future directions for a neglected phylum'.
Keywords: doubly uniparental inheritance; evolution; genome; mitochondria; mollusc.
Figures
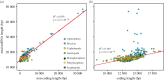

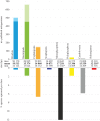
Similar articles
-
Mobilizing molluscan models and genomes in biology.Philos Trans R Soc Lond B Biol Sci. 2021 May 24;376(1825):20200163. doi: 10.1098/rstb.2020.0163. Epub 2021 Apr 5. Philos Trans R Soc Lond B Biol Sci. 2021. PMID: 33813892 Free PMC article.
-
Molluscan phylogenomics requires strategically selected genomes.Philos Trans R Soc Lond B Biol Sci. 2021 May 24;376(1825):20200161. doi: 10.1098/rstb.2020.0161. Epub 2021 Apr 5. Philos Trans R Soc Lond B Biol Sci. 2021. PMID: 33813889 Free PMC article.
-
Single individual structural variant detection uncovers widespread hemizygosity in molluscs.Philos Trans R Soc Lond B Biol Sci. 2021 May 24;376(1825):20200153. doi: 10.1098/rstb.2020.0153. Epub 2021 Apr 5. Philos Trans R Soc Lond B Biol Sci. 2021. PMID: 33813894 Free PMC article.
-
Potential of genomic technologies to improve disease resistance in molluscan aquaculture.Philos Trans R Soc Lond B Biol Sci. 2021 May 24;376(1825):20200168. doi: 10.1098/rstb.2020.0168. Epub 2021 Apr 5. Philos Trans R Soc Lond B Biol Sci. 2021. PMID: 33813884 Free PMC article. Review.
-
Doubly uniparental inheritance: two mitochondrial genomes, one precious model for organelle DNA inheritance and evolution.DNA Cell Biol. 2009 Feb;28(2):79-89. doi: 10.1089/dna.2008.0807. DNA Cell Biol. 2009. PMID: 19196051 Review.
Cited by
-
Genetic Insights into the Giant Keyhole Limpet (Megathura crenulata), an Eastern Pacific Coastal Endemic: Complete Mitogenome, Phylogenetics, Phylogeography, and Historical Demography.Genes (Basel). 2024 Oct 8;15(10):1303. doi: 10.3390/genes15101303. Genes (Basel). 2024. PMID: 39457427 Free PMC article.
-
High Connectivity at Abyssal Depths: Genomic and Proteomic Insights Into Population Structure of the Pan-Atlantic Deep-Sea Bivalve Ledella ultima (E. A. Smith, 1885).Ecol Evol. 2025 Aug 8;15(8):e71903. doi: 10.1002/ece3.71903. eCollection 2025 Aug. Ecol Evol. 2025. PMID: 40785988 Free PMC article.
-
ORFans in Mitochondrial Genomes of Marine Polychaete Polydora.Genome Biol Evol. 2023 Dec 1;15(12):evad219. doi: 10.1093/gbe/evad219. Genome Biol Evol. 2023. PMID: 38019573 Free PMC article.
-
The complete mitochondrial genome data of Pholas orientalis (Gmelin, 1791) from Malaysia.Data Brief. 2024 Jun 2;55:110581. doi: 10.1016/j.dib.2024.110581. eCollection 2024 Aug. Data Brief. 2024. PMID: 38966661 Free PMC article.
-
Comparative mitochondrial genome analysis provides new insights into the classification of Modiolinae.Mol Biol Rep. 2024 Jul 18;51(1):823. doi: 10.1007/s11033-024-09767-0. Mol Biol Rep. 2024. PMID: 39023631
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

