Liquid-like protein interactions catalyse assembly of endocytic vesicles
- PMID: 33820972
- PMCID: PMC8035231
- DOI: 10.1038/s41556-021-00646-5
Liquid-like protein interactions catalyse assembly of endocytic vesicles
Abstract
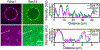
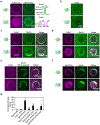
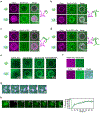
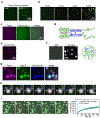
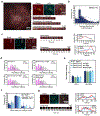
During clathrin-mediated endocytosis, dozens of proteins assemble into an interconnected network at the plasma membrane. As initiators of endocytosis, Eps15 and Fcho1/2 concentrate downstream components, while permitting dynamic rearrangement within the budding vesicle. How do initiator proteins meet these competing demands? Here we show that Eps15 and Fcho1/2 rely on weak, liquid-like interactions to catalyse endocytosis. In vitro, these weak interactions promote the assembly of protein droplets with liquid-like properties. To probe the physiological role of these liquid-like networks, we tuned the strength of initiator protein assembly in real time using light-inducible oligomerization of Eps15. Low light levels drove liquid-like assemblies, restoring normal rates of endocytosis in mammalian Eps15-knockout cells. By contrast, initiator proteins formed solid-like assemblies upon exposure to higher light levels, which stalled vesicle budding, probably owing to insufficient molecular rearrangement. These findings suggest that liquid-like assembly of initiator proteins provides an optimal catalytic platform for endocytosis.
Conflict of interest statement
Competing Interests
The authors declare no competing interests.
Figures







Comment in
-
Liquid-like protein assemblies initiate endocytosis.Nat Cell Biol. 2021 Apr;23(4):301-302. doi: 10.1038/s41556-021-00665-2. Nat Cell Biol. 2021. PMID: 33820974 No abstract available.
Similar articles
-
Pan1p, yeast eps15, functions as a multivalent adaptor that coordinates protein-protein interactions essential for endocytosis.J Cell Biol. 1998 Apr 6;141(1):71-84. doi: 10.1083/jcb.141.1.71. J Cell Biol. 1998. PMID: 9531549 Free PMC article.
-
Transient Fcho1/2⋅Eps15/R⋅AP-2 Nanoclusters Prime the AP-2 Clathrin Adaptor for Cargo Binding.Dev Cell. 2016 Jun 6;37(5):428-43. doi: 10.1016/j.devcel.2016.05.003. Epub 2016 May 26. Dev Cell. 2016. PMID: 27237791 Free PMC article.
-
Eps15 homology domain-NPF motif interactions regulate clathrin coat assembly during synaptic vesicle recycling.J Biol Chem. 2003 Aug 29;278(35):33583-92. doi: 10.1074/jbc.M304346200. Epub 2003 Jun 14. J Biol Chem. 2003. PMID: 12807910
-
Membrane remodeling in clathrin-mediated endocytosis.J Cell Sci. 2018 Sep 3;131(17):jcs216812. doi: 10.1242/jcs.216812. J Cell Sci. 2018. PMID: 30177505 Review.
-
Biophysics of endocytic vesicle formation: A focus on liquid-liquid phase separation.Curr Opin Cell Biol. 2022 Apr;75:102068. doi: 10.1016/j.ceb.2022.02.002. Epub 2022 Mar 10. Curr Opin Cell Biol. 2022. PMID: 35279562 Review.
Cited by
-
Dipping contacts - a novel type of contact site at the interface between membraneless organelles and membranes.J Cell Sci. 2023 Dec 15;136(24):jcs261413. doi: 10.1242/jcs.261413. Epub 2023 Dec 27. J Cell Sci. 2023. PMID: 38149872 Free PMC article.
-
Multiple polarity kinases inhibit phase separation of F-BAR protein Cdc15 and antagonize cytokinetic ring assembly in fission yeast.Elife. 2023 Feb 7;12:e83062. doi: 10.7554/eLife.83062. Elife. 2023. PMID: 36749320 Free PMC article.
-
Selective Endocytic Uptake of Targeted Liposomes Occurs within a Narrow Range of Liposome Diameters.ACS Appl Mater Interfaces. 2023 Nov 1;15(43):49988-50001. doi: 10.1021/acsami.3c09399. Epub 2023 Oct 20. ACS Appl Mater Interfaces. 2023. PMID: 37862704 Free PMC article.
-
Orchestrating vesicular and nonvesicular membrane dynamics by intrinsically disordered proteins.EMBO Rep. 2023 Nov 6;24(11):e57758. doi: 10.15252/embr.202357758. Epub 2023 Sep 8. EMBO Rep. 2023. PMID: 37680133 Free PMC article. Review.
-
Synapsin condensation controls synaptic vesicle sequestering and dynamics.Nat Commun. 2023 Oct 23;14(1):6730. doi: 10.1038/s41467-023-42372-6. Nat Commun. 2023. PMID: 37872159 Free PMC article.
References
Methods References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

