FOXO1 controls protein synthesis and transcript abundance of mutant polyglutamine proteins, preventing protein aggregation
- PMID: 33822053
- PMCID: PMC8170844
- DOI: 10.1093/hmg/ddab095
FOXO1 controls protein synthesis and transcript abundance of mutant polyglutamine proteins, preventing protein aggregation
Abstract
FOXO1, a transcription factor downstream of the insulin/insulin like growth factor axis, has been linked to protein degradation. Elevated expression of FOXO orthologs can also prevent the aggregation of cytosine adenine guanine (CAG)-repeat disease causing polyglutamine (polyQ) proteins but whether FOXO1 targets mutant proteins for degradation is unclear. Here, we show that increased expression of FOXO1 prevents toxic polyQ aggregation in human cells while reducing FOXO1 levels has the opposite effect and accelerates it. Although FOXO1 indeed stimulates autophagy, its effect on polyQ aggregation is independent of autophagy, ubiquitin-proteasome system (UPS) mediated protein degradation and is not due to a change in mutant polyQ protein turnover. Instead, FOXO1 specifically downregulates protein synthesis rates from expanded pathogenic CAG repeat transcripts. FOXO1 orchestrates a change in the composition of proteins that occupy mutant expanded CAG transcripts, including the recruitment of IGF2BP3. This mRNA binding protein enables a FOXO1 driven decrease in pathogenic expanded CAG transcript- and protein levels, thereby reducing the initiation of amyloidogenesis. Our data thus demonstrate that FOXO1 not only preserves protein homeostasis at multiple levels, but also reduces the accumulation of aberrant RNA species that may co-contribute to the toxicity in CAG-repeat diseases.
© The Author(s) 2021. Published by Oxford University Press. All rights reserved. For Permissions, please email: journals.permissions@oup.com.
Figures




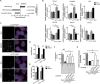
References
-
- Balch, W.E., Morimoto, R.I., Dillin, A. et al. (2008) Adapting proteostasis for disease intervention. Science (80-.)., 319, 916–919. - PubMed
-
- Kampinga, H.H. and Bergink, S. (2016) Heat shock proteins as potential targets for protective strategies in neurodegeneration. Lancet Neurol., 15, 748–759. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

