A genome sequence from a modern human skull over 45,000 years old from Zlatý kůň in Czechia
- PMID: 33828249
- PMCID: PMC8175239
- DOI: 10.1038/s41559-021-01443-x
A genome sequence from a modern human skull over 45,000 years old from Zlatý kůň in Czechia
Abstract
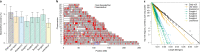
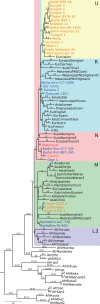
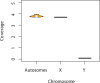
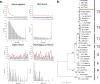
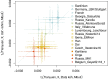
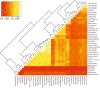
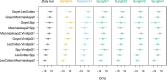
Modern humans expanded into Eurasia more than 40,000 years ago following their dispersal out of Africa. These Eurasians carried ~2-3% Neanderthal ancestry in their genomes, originating from admixture with Neanderthals that took place sometime between 50,000 and 60,000 years ago, probably in the Middle East. In Europe, the modern human expansion preceded the disappearance of Neanderthals from the fossil record by 3,000-5,000 years. The genetic makeup of the first Europeans who colonized the continent more than 40,000 years ago remains poorly understood since few specimens have been studied. Here, we analyse a genome generated from the skull of a female individual from Zlatý kůň, Czechia. We found that she belonged to a population that appears to have contributed genetically neither to later Europeans nor to Asians. Her genome carries ~3% Neanderthal ancestry, similar to those of other Upper Palaeolithic hunter-gatherers. However, the lengths of the Neanderthal segments are longer than those observed in the currently oldest modern human genome of the ~45,000-year-old Ust'-Ishim individual from Siberia, suggesting that this individual from Zlatý kůň is one of the earliest Eurasian inhabitants following the expansion out of Africa.
Conflict of interest statement
The authors declare no competing interests.
Figures






Comment in
-
Neanderthal assimilation?Nat Ecol Evol. 2021 Jun;5(6):711-712. doi: 10.1038/s41559-021-01421-3. Nat Ecol Evol. 2021. PMID: 33828250 No abstract available.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

