Grass Carp Reovirus VP35 Degrades MAVS Through the Autophagy Pathway to Inhibit Fish Interferon Production
- PMID: 33833752
- PMCID: PMC8021864
- DOI: 10.3389/fimmu.2021.613145
Grass Carp Reovirus VP35 Degrades MAVS Through the Autophagy Pathway to Inhibit Fish Interferon Production
Abstract
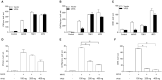
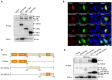
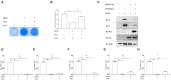
Fish interferon (IFN) is a crucial cytokine for a host to resist external pathogens, conferring cells with antiviral capacity. Meanwhile, grass carp reovirus (GCRV) is a strong pathogen that causes high mortality in grass carp. Therefore, it is necessary to study the strategy used by GCRV to evade the cellular IFN response. In this study, we found that GCRV 35-kDa protein (VP35) inhibited the host IFN production by degrading mitochondrial antiviral signaling (MAVS) protein through the autophagy pathway. First, the overexpression of VP35 inhibited the IFN activation induced by polyinosinic-polycytidylic acid (poly I:C) and MAVS, and the expression of downstream IFN-stimulated genes (ISGs) was also decreased by using VP35 under the stimulation. Second, VP35 interacted with MAVS; the experiments of truncated mutants of MAVS demonstrated that the caspase recruitment domain (CARD) and proline-rich (PRO) domains of MAVS were not necessary for this binding. Then, MAVS was degraded by using VP35 in a dose-dependent manner, and 3-MA (the autophagy pathway inhibitor) significantly blocked the degradation, meaning that MAVS was degraded by using VP35 in the autophagy pathway. The result of MAVS degradation suggested that the antiviral capacity of MAVS was remarkably depressed when interrupted by VP35. Finally, in the host cells, VP35 reduced ifn transcription and made the cells vulnerable to virus infection. In conclusion, our results reveal that GCRV VP35 impairs the host IFN response by degrading MAVS through the autophagy pathway, supplying evidence of a fish virus immune evasion strategy.
Keywords: GCRV; MAVS; VP35; immune evasion; interferon.
Copyright © 2021 Lu, Zhang, Li, Zhou, Jiang, Chen, Zhang and Li.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




References
-
- Yang Y, Peng ZQ, Li H, Tan SW, Yu HY, Yu H. Epidemiological survey of grass carp (Ctenopharyngodon idella) reovirus in south china, and genetic variations of VP6 gene. Isr J Aquac. (2017) 69:1390–99. 10.46989/001c.21049 - DOI
-
- Zeng WW, Yao W, Wang YY, Li YY, Bermann SM, Ren Y, et al. Molecular detection of genotype II grass carp reovirus based on nucleic acid sequence-based amplification combined with enzyme-linked immunosorbent assay (NASBA-ELISA). J Virol Methods. (2017) 243:92–7. 10.1016/j.jviromet.2017.02.001 - DOI - PubMed
-
- Attoui H, Fang Q, Jaafar FM, Cantaloube JF, Biagini P, de Micco P, et al. Common evolutionary origin of aquareoviruses and orthoreoviruses revealed by genome characterization of Golden shiner reovirus, Grass carp reovirus, Striped bass reovirus and golden ide reovirus (genus Aquareovirus, family Reoviridae). J Gen Virol. (2002) 83:1941–51. 10.1099/0022-1317-83-8-1941 - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

