Legume-microbiome interactions unlock mineral nutrients in regrowing tropical forests
- PMID: 33836596
- PMCID: PMC7980381
- DOI: 10.1073/pnas.2022241118
Legume-microbiome interactions unlock mineral nutrients in regrowing tropical forests
Abstract
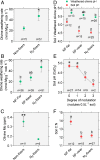
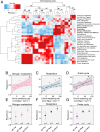
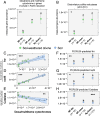
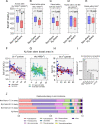
Legume trees form an abundant and functionally important component of tropical forests worldwide with N2-fixing symbioses linked to enhanced growth and recruitment in early secondary succession. However, it remains unclear how N2-fixers meet the high demands for inorganic nutrients imposed by rapid biomass accumulation on nutrient-poor tropical soils. Here, we show that N2-fixing trees in secondary Neotropical forests triggered twofold higher in situ weathering of fresh primary silicates compared to non-N2-fixing trees and induced locally enhanced nutrient cycling by the soil microbiome community. Shotgun metagenomic data from weathered minerals support the role of enhanced nitrogen and carbon cycling in increasing acidity and weathering. Metagenomic and marker gene analyses further revealed increased microbial potential beneath N2-fixers for anaerobic iron reduction, a process regulating the pool of phosphorus bound to iron-bearing soil minerals. We find that the Fe(III)-reducing gene pool in soil is dominated by acidophilic Acidobacteria, including a highly abundant genus of previously undescribed bacteria, Candidatus Acidoferrum, genus novus. The resulting dependence of the Fe-cycling gene pool to pH determines the high iron-reducing potential encoded in the metagenome of the more acidic soils of N2-fixers and their nonfixing neighbors. We infer that by promoting the activities of a specialized local microbiome through changes in soil pH and C:N ratios, N2-fixing trees can influence the wider biogeochemical functioning of tropical forest ecosystems in a manner that enhances their ability to assimilate and store atmospheric carbon.
Keywords: Acidobacteria; N2-fixing legume trees; metagenomics; mineral weathering; tropical forest.
Copyright © 2021 the Author(s). Published by PNAS.
Conflict of interest statement
The authors declare no competing interest.
Figures

References
-
- ter Steege H., et al., Continental-scale patterns of canopy tree composition and function across Amazonia. Nature 443, 444–447 (2006). - PubMed
-
- Hedin L. O., Brookshire E. N. J., Menge D. N. L., Barron A. R., The nitrogen paradox in tropical forest ecosystems. Annu. Rev. Ecol. Evol. Syst. 40, 613–635 (2009).
-
- Sprent J. I., Ardley J., James E. K., Biogeography of nodulated legumes and their nitrogen-fixing symbionts. New Phytol. 215, 40–56 (2017). - PubMed
-
- Batterman S. A., et al., Key role of symbiotic dinitrogen fixation in tropical forest secondary succession. Nature 502, 224–227 (2013). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

