Gene expression and immune infiltration in melanoma patients with different mutation burden
- PMID: 33836680
- PMCID: PMC8034108
- DOI: 10.1186/s12885-021-08083-1
Gene expression and immune infiltration in melanoma patients with different mutation burden
Abstract
Background: Immunotherapy is a vital component in cancer treatment. However, due to the complex genetic bases of cancer, a clear prediction index for efficacy has not been established. Tumor mutation burden (TMB) is one of the essential factors that affect immunotherapeutic efficacies, but it has not been determined whether the mutation is associated with the survival of Skin Cutaneous Melanoma (SKCM) patients. This study aimed at evaluating the correlation between TMB and immune infiltration.
Methods: Somatic mutation profiles (n = 467), transcriptome data (n = 471), and their clinical information (n = 447) of all SKCM samples were downloaded from The Cancer Genome Atlas (TCGA) database. For each sample, TMB was calculated as the number of variants per megabase. Based on K-M survival analysis, they were allocated into the high-TMB and low-TMB groups (the optimal cutoff was determined by the 'surv_cutpoint' algorithm of survival R package). Then, Gene ontology (GO) and Gene Set Enrichment Analyses (GSEA) were performed, with immune-associated biological pathways found to be significantly enriched in the low-TMB group. Therefore, immune genes that were differentially expressed between the two groups were evaluated in Cox regression to determine their prognostic values, and a four-gene TMB immune prognostic model (TMB-IP) was constructed.
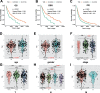
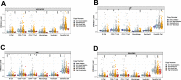
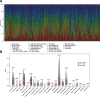
Results: Elevated TMB levels were associated with better survival outcomes in SKCM patients. Based on the cutoff value in OS analysis, they were divided into high-TMB and low-TMB groups. GSEA revealed that the low-TMB group was associated with immunity while intersection analysis revealed that there were 38 differentially expressed immune-related genes between the two groups. Four TMB-associated immune genes were used to construct a TMB-IP model. The AUC of the ROC curve of this model reached a maximum of 0.75 (95%CI, 0.66-0.85) for OS outcomes. Validation in each clinical subgroup confirmed the efficacy of the model to distinguish between high and low TMB-IP score patients.
Conclusions: In SKCM patients, low TMB was associated with worse survival outcomes and enriched immune-associated pathways. The four TMB-associated immune genes model can effectively distinguish between high and low-risk patients.
Keywords: ICB; SKCM; Survival analysis; TCGA; TMB.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures


Similar articles
-
Multi-omics analysis of tumor mutation burden combined with immune infiltrates in melanoma.Clin Chim Acta. 2020 Dec;511:306-318. doi: 10.1016/j.cca.2020.10.030. Epub 2020 Oct 24. Clin Chim Acta. 2020. PMID: 33164879
-
The prognostic value of TMB and the relationship between TMB and immune infiltration in head and neck squamous cell carcinoma: A gene expression-based study.Oral Oncol. 2020 Nov;110:104943. doi: 10.1016/j.oraloncology.2020.104943. Epub 2020 Sep 9. Oral Oncol. 2020. PMID: 32919362
-
Tumor Mutation Burden, Immune Cell Infiltration, and Construction of Immune-Related Genes Prognostic Model in Head and Neck Cancer.Int J Med Sci. 2021 Jan 1;18(1):226-238. doi: 10.7150/ijms.51064. eCollection 2021. Int J Med Sci. 2021. PMID: 33390791 Free PMC article.
-
A Pan-Cancer Analysis Reveals the Prognostic and Immunotherapeutic Value of Stanniocalcin-2 (STC2).Front Genet. 2022 Jul 22;13:927046. doi: 10.3389/fgene.2022.927046. eCollection 2022. Front Genet. 2022. PMID: 35937984 Free PMC article. Review.
-
TMBserval: a statistical explainable learning model reveals weighted tumor mutation burden better categorizing therapeutic benefits.Front Immunol. 2023 May 10;14:1151755. doi: 10.3389/fimmu.2023.1151755. eCollection 2023. Front Immunol. 2023. PMID: 37234148 Free PMC article.
Cited by
-
Therapeutic implications of endoplasmic reticulum stress gene CCL3 in cervical squamous cell carcinoma.Cell Biol Toxicol. 2025 Feb 20;41(1):47. doi: 10.1007/s10565-024-09949-3. Cell Biol Toxicol. 2025. PMID: 39976849 Free PMC article.
-
Current State of Melanoma Therapy and Next Steps: Battling Therapeutic Resistance.Cancers (Basel). 2024 Apr 19;16(8):1571. doi: 10.3390/cancers16081571. Cancers (Basel). 2024. PMID: 38672652 Free PMC article. Review.
-
Predictive Prognostic Model for Hepatocellular Carcinoma Based on Seven Genes Participating in Arachidonic Acid Metabolism.Cancer Med. 2024 Nov;13(22):e70284. doi: 10.1002/cam4.70284. Cancer Med. 2024. PMID: 39540710 Free PMC article.
-
S1P-S1PR3-RAS promotes the progression of S1PR3hi TAL1+ T-cell acute lymphoblastic leukemia that can be effectively inhibited by an S1PR3 antagonist.Leukemia. 2023 Oct;37(10):1982-1993. doi: 10.1038/s41375-023-02000-0. Epub 2023 Aug 17. Leukemia. 2023. PMID: 37591940
-
Comprehensive Pan-Cancer Mutation Density Patterns in Enhancer RNA.Int J Mol Sci. 2023 Dec 30;25(1):534. doi: 10.3390/ijms25010534. Int J Mol Sci. 2023. PMID: 38203707 Free PMC article.
References
-
- De La Cruz MM, Abdul Z, Shariff Z. The impact of a skin cancer diagnosis on travel insurance: a sun worshipper's dilemma. Clin Exp Dermatol. 2020. - PubMed
-
- Siegel JA, Yudkin JS, Craker K, Hwang A, Libby T. Uncapping the bottle: a proposal to allow full-sized sunscreens in carry-on luggage to promote sun protection and prevent skin cancer. J Am Acad Dermatol. 2020. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

