LINC01123 enhances osteosarcoma cell growth by activating the Hedgehog pathway via the miR-516b-5p/Gli1 axis
- PMID: 33837611
- PMCID: PMC8177773
- DOI: 10.1111/cas.14913
LINC01123 enhances osteosarcoma cell growth by activating the Hedgehog pathway via the miR-516b-5p/Gli1 axis
Abstract
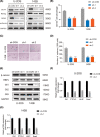
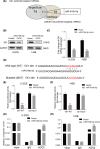
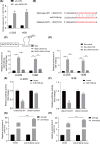
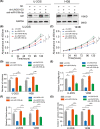
The lncRNA LINC01123 has been reported to act as an oncogene in many human cancers. Nevertheless, the function and underlying mechanism of LINC01123 in osteosarcoma (OS) remain unclear. This study aimed to explore the roles and mechanisms of LINC01123 in OS progression. In this study, the expression of LINC01123 was significantly upregulated in OS cell lines than in a human osteoblast cell line. Furthermore, in vitro and in vivo experiments confirmed that knockdown of LINC01123 suppressed cell progression. Mechanistically, LINC01123 acted as a competing endogenous RNA by sponging miR-516b-5p, thus, increasing Gli1 expression by directly targeting its 3'UTR. Taken together, LINC01123 enhances OS proliferation and metastasis via the miR-516b-5p/Gli1 axis. Therefore, LINC01123 may be a potential therapeutic target for OS treatment.
Keywords: Gli1; LINC01123; metastasis; osteosarcoma; proliferation.
© 2021 The Authors. Cancer Science published by John Wiley & Sons Australia, Ltd on behalf of Japanese Cancer Association.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures



Similar articles
-
Long non-coding RNA XIST promotes osteosarcoma progression by targeting YAP via miR-195-5p.J Cell Biochem. 2018 Jul;119(7):5646-5656. doi: 10.1002/jcb.26743. Epub 2018 Mar 25. J Cell Biochem. 2018. Retraction in: J Cell Biochem. 2021 Apr;122(3-4):485. doi: 10.1002/jcb.29872. PMID: 29384226 Retracted.
-
NEAT1 induces osteosarcoma development by modulating the miR-339-5p/TGF-β1 pathway.J Cell Physiol. 2019 Apr;234(4):5097-5105. doi: 10.1002/jcp.27313. Epub 2018 Sep 10. J Cell Physiol. 2019. PMID: 30203547
-
lncRNA nuclear-enriched abundant transcript 1 promotes cell proliferation and invasion by targeting miR-186-5p/HIF-1α in osteosarcoma.J Cell Biochem. 2019 Apr;120(4):6502-6514. doi: 10.1002/jcb.27941. Epub 2018 Nov 28. J Cell Biochem. 2019. PMID: 30485482
-
The critical roles of lncRNAs in the development of osteosarcoma.Biomed Pharmacother. 2021 Mar;135:111217. doi: 10.1016/j.biopha.2021.111217. Epub 2021 Feb 1. Biomed Pharmacother. 2021. PMID: 33433358 Review.
-
Research progress on the role of lncRNA, circular RNA, and microRNA networks in regulating ferroptosis in osteosarcoma.Biomed Pharmacother. 2024 Jul;176:116924. doi: 10.1016/j.biopha.2024.116924. Epub 2024 Jun 14. Biomed Pharmacother. 2024. PMID: 38876052 Review.
Cited by
-
Noncoding RNAs related to the hedgehog pathway in cancer: clinical implications and future perspectives.Mol Cancer. 2022 May 17;21(1):115. doi: 10.1186/s12943-022-01591-z. Mol Cancer. 2022. PMID: 35581586 Free PMC article. Review.
-
Unfolding antifungals: as a new foe to pancreatic ductal adenocarcinoma-a mini-review.Mol Biol Rep. 2021 Mar;48(3):2945-2956. doi: 10.1007/s11033-021-06318-9. Epub 2021 Apr 1. Mol Biol Rep. 2021. PMID: 33796989 Review.
-
LncRNA LINC01197 inhibited the formation of calcium oxalate-induced kidney stones by regulating miR-516b-5p/SIRT3/FOXO1 signaling pathway.Cell Tissue Res. 2023 May;392(2):553-563. doi: 10.1007/s00441-022-03734-6. Epub 2023 Jan 23. Cell Tissue Res. 2023. PMID: 36688989
-
Immune-Related LncRNAs Affect the Prognosis of Osteosarcoma, Which Are Related to the Tumor Immune Microenvironment.Front Cell Dev Biol. 2021 Oct 7;9:731311. doi: 10.3389/fcell.2021.731311. eCollection 2021. Front Cell Dev Biol. 2021. PMID: 34692688 Free PMC article.
-
CircRNA OXCT1 promotes the malignant progression and glutamine metabolism of non-small cell lung cancer by absorbing miR-516b-5p and upregulating SLC1A5.Cell Cycle. 2023 May;22(10):1182-1195. doi: 10.1080/15384101.2022.2071565. Epub 2023 Mar 2. Cell Cycle. 2023. PMID: 35482822 Free PMC article.
References
-
- Ritter J, Bielack Ss. Osteosarcoma. Ann Oncol. 2010;7:vii320–vii325. - PubMed
-
- Meazza C, Scanagatta P. Metastatic osteosarcoma: a challenging multidisciplinary treatment. Expert Rev Anticancer Ther. 2016;16:543‐556. - PubMed
-
- Quinn JJ, Chang HY. Unique features of long non‐coding RNA biogenesis and function. Nat Rev Genet. 2016;17:47‐62. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

