CircRNA_100395 Carried by Exosomes From Adipose-Derived Mesenchymal Stem Cells Inhibits the Malignant Transformation of Non-Small Cell Lung Carcinoma Through the miR-141-3p-LATS2 Axis
- PMID: 33842488
- PMCID: PMC8027360
- DOI: 10.3389/fcell.2021.663147
CircRNA_100395 Carried by Exosomes From Adipose-Derived Mesenchymal Stem Cells Inhibits the Malignant Transformation of Non-Small Cell Lung Carcinoma Through the miR-141-3p-LATS2 Axis
Abstract
Objective: The specific purpose of this study is to investigate the impact exosomes from adipose-derived mesenchymal stem cell (AMSC) has on non-small cell lung carcinoma (NSCLC) and the relative applications.
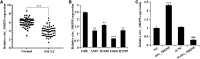
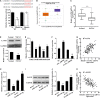
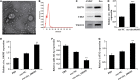
Methods: circ_100395, miR-141-3p, and LATS2 were expressed and detected in NSCLC and paracancerous tissues as well as NSCLC cell lines. Pearson correlation analysis, Dual-Luciferase Reporter Assay and RNA pull-down assay were used to validate their expression and interaction, respectively. After isolation and culture of AMSCs, exosomes were extracted and identified. EdU, epithelial-mesenchymal transition (EMT), and cell colony formation assay were used to distinguish the biological activity of the cells. Expression Hippo/YAP signalling pathway-related proteins were measured by western blotting. Subsequently, tumour volume and weight were confirmed based on xenograft nude mice models, Ki-67 and LATS2 expression was observed by immunohistochemistry.
Results: circ_100395 was lowly expressed in NSCLC tissues or cells. The negative correlations and interactions were confirmed between circ_100395 and miR-141-3p, miR-141-3p, and LATS2. AMSC-derived exosomes with overexpression of circ_100395 (exo-circ_100395) significantly inhibited the biological activity as well as EMT of H1650 cells and Hippo/YAP signalling pathway activity. In addition, exo-circ_100395 markedly reduced tumour volume and weight as well as Ki-67 and LASP1 expression in vivo. However, overexpressed miR-141-3p or knocked down LATS2 alleviated the above effects.
Conclusion: Exo-circ_100395 can increase LATS2 expression by sponging miR-141-3p to regulate Hippo/YAP signalling pathway, thereby inhibiting NSCLC malignant transformation.
Keywords: Hippo/YAP signalling pathway; circRNA_100395; exosomes from adipose-derived mesenchymal stem cells; miR-141-3p; non-small cell lung carcinoma (NSCLC).
Copyright © 2021 Zhang, Cao, Lv and Mou.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




Similar articles
-
circRNA_0000140 suppresses oral squamous cell carcinoma growth and metastasis by targeting miR-31 to inhibit Hippo signaling pathway.Cell Death Dis. 2020 Feb 10;11(2):112. doi: 10.1038/s41419-020-2273-y. Cell Death Dis. 2020. PMID: 32041942 Free PMC article.
-
N6-Methyladenosine Reader YTHDF2 Enhances Non-Small-Cell Lung Cancer Cell Proliferation and Metastasis through Mediating circ_SFMBT2 Degradation.Contrast Media Mol Imaging. 2022 Jul 16;2022:1087622. doi: 10.1155/2022/1087622. eCollection 2022. Contrast Media Mol Imaging. 2022. PMID: 35924072 Free PMC article.
-
Exosomal Circ-MEMO1 Promotes the Progression and Aerobic Glycolysis of Non-small Cell Lung Cancer Through Targeting MiR-101-3p/KRAS Axis.Front Genet. 2020 Aug 28;11:962. doi: 10.3389/fgene.2020.00962. eCollection 2020. Front Genet. 2020. PMID: 33005174 Free PMC article.
-
MiR-199a-modified exosomes from adipose tissue-derived mesenchymal stem cells improve hepatocellular carcinoma chemosensitivity through mTOR pathway.J Exp Clin Cancer Res. 2020 Jan 2;39(1):4. doi: 10.1186/s13046-019-1512-5. J Exp Clin Cancer Res. 2020. PMID: 31898515 Free PMC article.
-
Downregulation of microRNA-224-3p Hampers Retinoblastoma Progression via Activation of the Hippo-YAP Signaling Pathway by Increasing LATS2.Invest Ophthalmol Vis Sci. 2020 Mar 9;61(3):32. doi: 10.1167/iovs.61.3.32. Invest Ophthalmol Vis Sci. 2020. PMID: 32186675 Free PMC article.
Cited by
-
The interplay between noncoding RNA and YAP/TAZ signaling in cancers: molecular functions and mechanisms.J Exp Clin Cancer Res. 2022 Jun 14;41(1):202. doi: 10.1186/s13046-022-02403-4. J Exp Clin Cancer Res. 2022. PMID: 35701841 Free PMC article. Review.
-
Large tumor suppressor 2 is a prognostic biomarker and correlated with immune infiltrates in colorectal cancer.Bioengineered. 2021 Dec;12(2):11648-11661. doi: 10.1080/21655979.2021.1996513. Bioengineered. 2021. PMID: 34699318 Free PMC article.
-
Exosome theranostics: Comparative analysis of P body and exosome proteins and their mutations for clinical applications.Funct Integr Genomics. 2024 Jul 12;24(4):124. doi: 10.1007/s10142-024-01404-0. Funct Integr Genomics. 2024. PMID: 38995459 Review.
-
Exosomal circRNAs: Emerging Players in Tumor Metastasis.Front Cell Dev Biol. 2021 Dec 8;9:786224. doi: 10.3389/fcell.2021.786224. eCollection 2021. Front Cell Dev Biol. 2021. PMID: 34957113 Free PMC article. Review.
-
CircRNA: A novel potential strategy to treat thyroid cancer (Review).Int J Mol Med. 2021 Nov;48(5):201. doi: 10.3892/ijmm.2021.5034. Epub 2021 Sep 16. Int J Mol Med. 2021. PMID: 34528697 Free PMC article. Review.
References
-
- Bourque D. K., Avila L., Peñaherrera M., von Dadelszen P., Robinson W. P. (2010). Decreased placental methylation at the H19/IGF2 imprinting control region is associated with normotensive intrauterine growth restriction but not preeclampsia. Placenta 31 197–202. 10.1016/j.placenta.2009.12.003 - DOI - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

